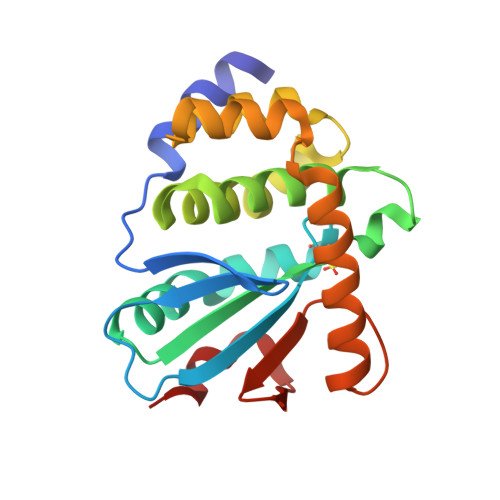
Structural Basis of a Thiol-Disulfide Oxidoreductase in the Hedgehog-Forming Actinobacterium Corynebacterium matruchotii.
Luong, T.T., Tirgar, R., Reardon-Robinson, M.E., Joachimiak, A., Osipiuk, J., Ton-That, H.(2018) J Bacteriol 200
- PubMed: 29440253
- DOI: https://doi.org/10.1128/JB.00783-17
- Primary Citation of Related Structures:
6BO0 - PubMed Abstract:
The actinobacterium Corynebacterium matruchotii has been implicated in nucleation of oral microbial consortia leading to biofilm formation. Due to the lack of genetic tools, little is known about basic cellular processes, including protein secretion and folding, in this organism. We report here a survey of the C. matruchotii genome, which encodes a large number of exported proteins containing paired cysteine residues, and identified an oxidoreductase that is highly homologous to the Corynebacterium diphtheriae thiol-disulfide oxidoreductase MdbA (MdbA Cd ). Crystallization studies uncovered that the 1.2-Å resolution structure of C. matruchotii MdbA (MdbA Cm ) possesses two conserved features found in actinobacterial MdbA enzymes, a thioredoxin-like fold and an extended α-helical domain. By reconstituting the disulfide bond-forming machine in vitro , we demonstrated that MdbA Cm catalyzes disulfide bond formation within the actinobacterial pilin FimA. A new gene deletion method supported that mdbA is essential in C. matruchotii Remarkably, heterologous expression of MdbA Cm in the C. diphtheriae Δ mdbA mutant rescued its known defects in cell growth and morphology, toxin production, and pilus assembly, and this thiol-disulfide oxidoreductase activity required the catalytic motif CXXC. Altogether, the results suggest that MdbA Cm is a major thiol-disulfide oxidoreductase, which likely mediates posttranslocational protein folding in C. matruchotii by a mechanism that is conserved in Actinobacteria IMPORTANCE The actinobacterium Corynebacterium matruchotii has been implicated in the development of oral biofilms or dental plaque; however, little is known about the basic cellular processes in this organism. We report here a high-resolution structure of a C. matruchotii oxidoreductase that is highly homologous to the Corynebacterium diphtheriae thiol-disulfide oxidoreductase MdbA. By biochemical analysis, we demonstrated that C. matruchotii MdbA catalyzes disulfide bond formation in vitro Furthermore, a new gene deletion method revealed that deletion of mdbA is lethal in C. matruchotii Remarkably, C. matruchotii MdbA can replace C. diphtheriae MdbA to maintain normal cell growth and morphology, toxin production, and pilus assembly. Overall, our studies support the hypothesis that C. matruchotii utilizes MdbA as a major oxidoreductase to catalyze oxidative protein folding.
Organizational Affiliation:
Department of Microbiology & Molecular Genetics, University of Texas Health Science Center, Houston, Texas, USA.