Molecular Insight into Recognition of the CGRPR Complex by Migraine Prevention Therapy Aimovig (Erenumab).
Garces, F., Mohr, C., Zhang, L., Huang, C.S., Chen, Q., King, C., Xu, C., Wang, Z.(2020) Cell Rep 30: 1714
- PubMed: 32049005
- DOI: https://doi.org/10.1016/j.celrep.2020.01.029
- Primary Citation of Related Structures:
6UMG, 6UMH, 6UMI, 6UMJ - PubMed Abstract:
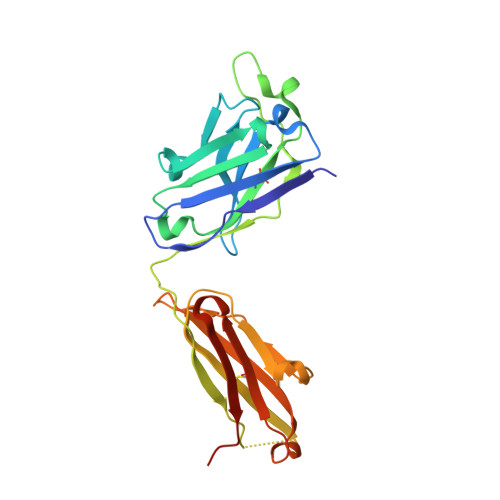
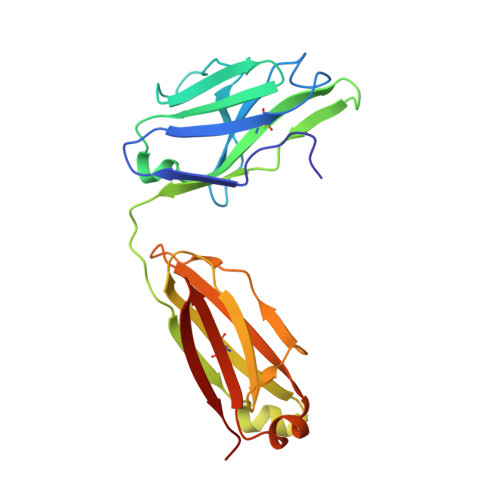
Calcitonin-gene-related peptide (CGRP) plays a key role in migraine pathophysiology. Aimovig (erenumab; erenumab-aooe in the United States) is the only US Food and Drug Administration (FDA)-approved monoclonal antibody (mAb) therapy against the CGRP receptor (CGRPR) for the prevention of migraine. Aimovig is also the first FDA-approved mAb against a G-protein-coupled receptor (GPCR). Here, we report the architecture and functional attributes of erenumab critical for its potent antagonism against CGRPR. The crystal structure of erenumab in complex with CGRPR reveals a direct ligand-blocking mechanism, enabled by a remarkable 21-residue-long complementary determining region (CDR)-H3 loop, which adopts a tyrosine-rich helix-turn tip and projects into the deep interface of the calcitonin receptor-like receptor (CLR) and RAMP1 subunits of CGRPR. Furthermore, erenumab engages with residues specific to CLR and RAMP1, providing the molecular basis for its exquisite selectivity. Such structural insights reveal the drug action mechanism of erenumab and shed light on developing antibody therapeutics targeting GPCRs.
Organizational Affiliation:
Department of Therapeutic Discovery, Amgen Research, Amgen, Thousand Oaks, CA 91320, USA.