Crystal structure of dihydropteroate synthase from Anaplasma phagocytophilum with bound 6-hydroxymethylpterin-monophosphate
Bolejack, M.J., Delker, S.L., Abendroth, J., Lorimer, D.D., Horanyi, P.S., Edwards, T.E.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Dihydropteroate synthase | 292 | Anaplasma phagocytophilum | Mutation(s): 0 Gene Names: WSQ_01200, folP EC: 2.5.1.15 | 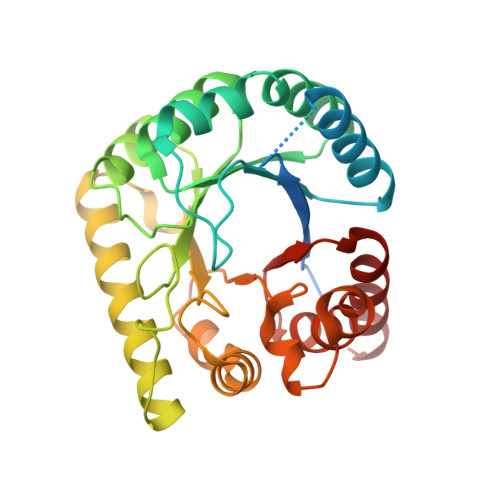 | |
UniProt | |||||
Find proteins for S5PWL8 (Anaplasma phagocytophilum str. JM) Explore S5PWL8 Go to UniProtKB: S5PWL8 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | S5PWL8 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PMM (Subject of Investigation/LOI) Query on PMM | C [auth A], G [auth B] | PTERIN-6-YL-METHYL-MONOPHOSPHATE C7 H8 N5 O5 P AJXFJEHKGGCFNM-UHFFFAOYSA-N |  | ||
| CIT Query on CIT | D [auth A] | CITRIC ACID C6 H8 O7 KRKNYBCHXYNGOX-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | H [auth B] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | E [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| NA Query on NA | F [auth A], I [auth B], J [auth B], K [auth B], L [auth B] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 48.83 | α = 90 |
| b = 81.48 | β = 102.67 |
| c = 71.17 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XDS | data reduction |
| XSCALE | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| MoRDa | phasing |