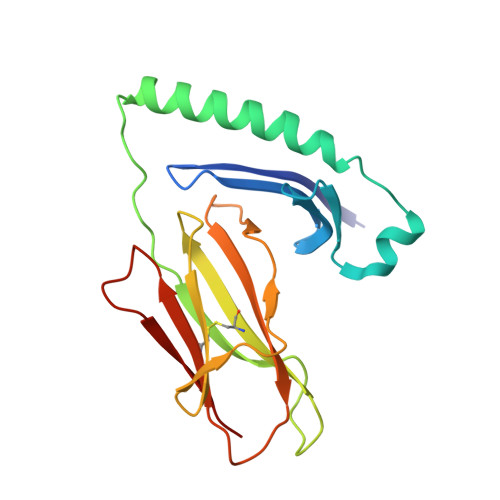
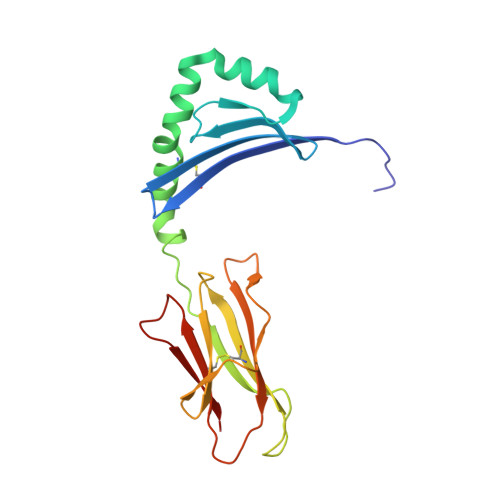
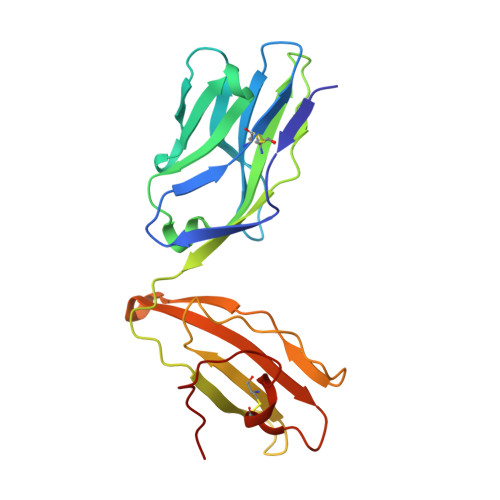
CD4+T Cells Recognize Conserved Influenza A Epitopes through Shared Patterns of V-Gene Usage and Complementary Biochemical Features.
Greenshields-Watson, A., Attaf, M., MacLachlan, B.J., Whalley, T., Rius, C., Wall, A., Lloyd, A., Hughes, H., Strange, K.E., Mason, G.H., Schauenburg, A.J., Hulin-Curtis, S.L., Geary, J., Chen, Y., Lauder, S.N., Smart, K., Vijaykrishna, D., Grau, M.L., Shugay, M., Andrews, R., Dolton, G., Rizkallah, P.J., Gallimore, A.M., Sewell, A.K., Godkin, A.J., Cole, D.K.(2020) Cell Rep 32: 107885-107885
- PubMed: 32668259
- DOI: https://doi.org/10.1016/j.celrep.2020.107885
- Primary Citation of Related Structures:
6QZA, 6QZC, 6QZD, 6R0E - PubMed Abstract:
T cell recognition of peptides presented by human leukocyte antigens (HLAs) is mediated by the highly variable T cell receptor (TCR). Despite this built-in TCR variability, individuals can mount immune responses against viral epitopes by using identical or highly related TCRs expressed on CD8 + T cells. Characterization of these TCRs has extended our understanding of the molecular mechanisms that govern the recognition of peptide-HLA. However, few examples exist for CD4 + T cells. Here, we investigate CD4 + T cell responses to the internal proteins of the influenza A virus that correlate with protective immunity. We identify five internal epitopes that are commonly recognized by CD4 + T cells in five HLA-DR1 + subjects and show conservation across viral strains and zoonotic reservoirs. TCR repertoire analysis demonstrates several shared gene usage biases underpinned by complementary biochemical features evident in a structural comparison. These epitopes are attractive targets for vaccination and other T cell therapies.
Organizational Affiliation:
Cardiff University, School of Medicine, Heath Park, Cardiff, UK.