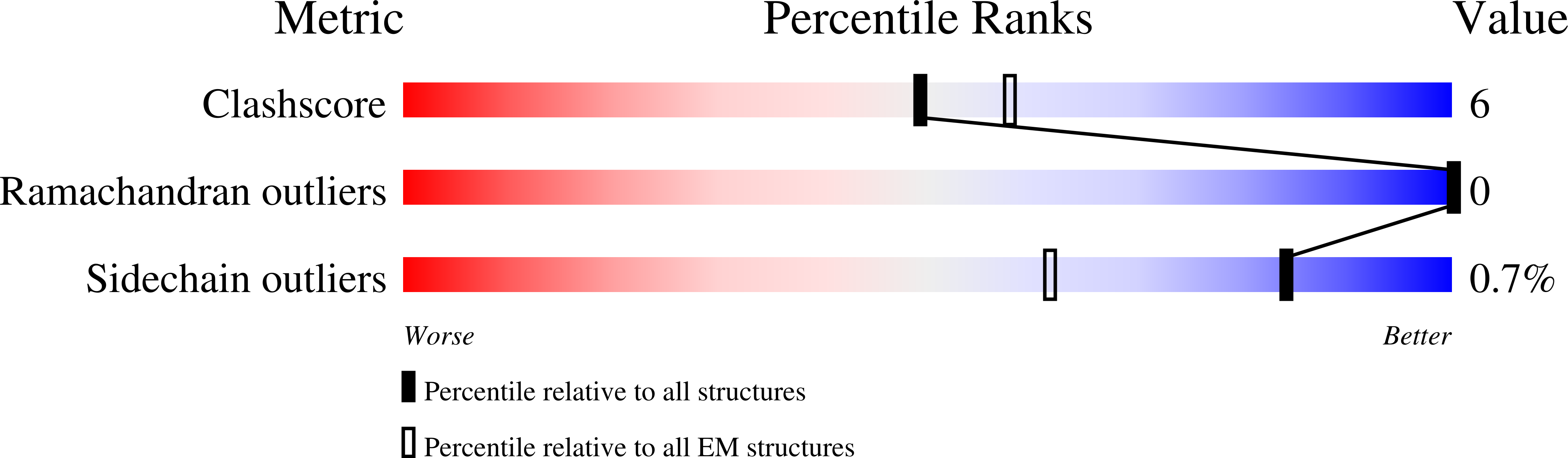Molecular basis of tRNA recognition by the Elongator complex.
Dauden, M.I., Jaciuk, M., Weis, F., Lin, T.Y., Kleindienst, C., Abbassi, N.E.H., Khatter, H., Krutyholowa, R., Breunig, K.D., Kosinski, J., Muller, C.W., Glatt, S.(2019) Sci Adv 5: eaaw2326-eaaw2326
- PubMed: 31309145
- DOI: https://doi.org/10.1126/sciadv.aaw2326
- Primary Citation of Related Structures:
6QK7 - PubMed Abstract:
The highly conserved Elongator complex modifies transfer RNAs (tRNAs) in their wobble base position, thereby regulating protein synthesis and ensuring proteome stability. The precise mechanisms of tRNA recognition and its modification reaction remain elusive. Here, we show cryo-electron microscopy structures of the catalytic subcomplex of Elongator and its tRNA-bound state at resolutions of 3.3 and 4.4 Å. The structures resolve details of the catalytic site, including the substrate tRNA, the iron-sulfur cluster, and a SAM molecule, which are all validated by mutational analyses in vitro and in vivo. tRNA binding induces conformational rearrangements, which precisely position the targeted anticodon base in the active site. Our results provide the molecular basis for substrate recognition of Elongator, essential to understand its cellular function and role in neurodegenerative diseases and cancer.
Organizational Affiliation:
European Molecular Biology Laboratory, Structural and Computational Biology Unit, Heidelberg, Germany.