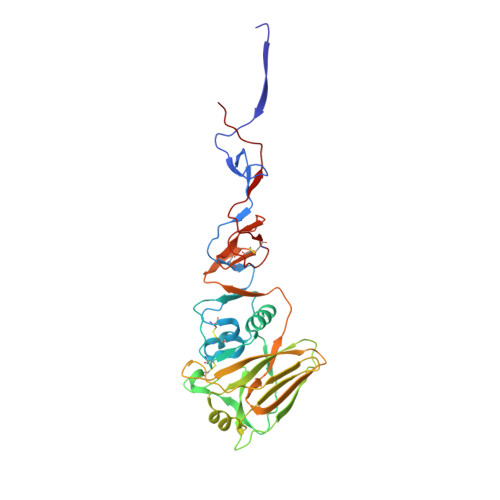
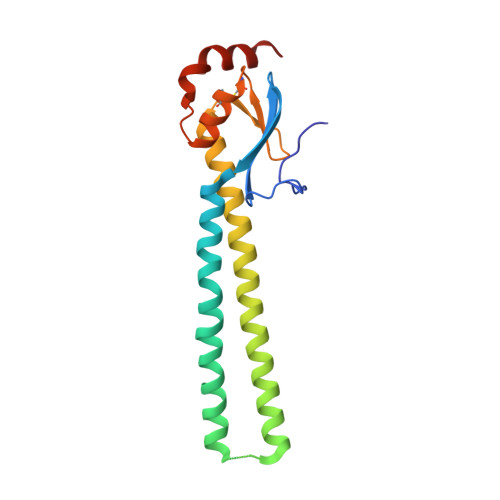
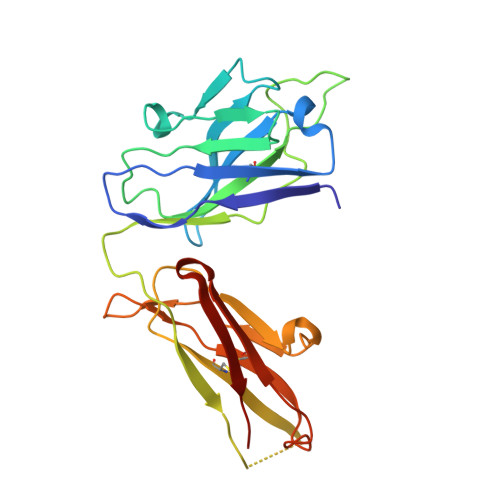
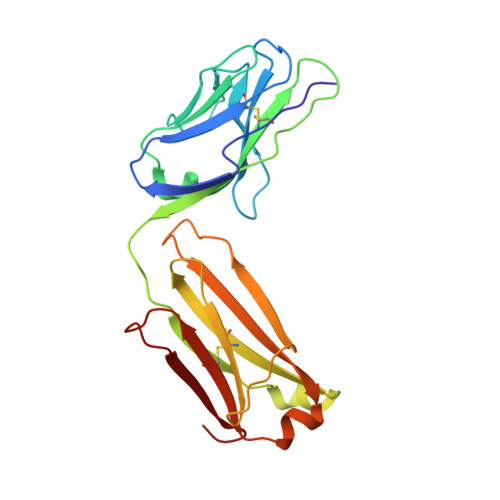
Prolonged evolution of the memory B cell response induced by a replicating adenovirus-influenza H5 vaccine.
Matsuda, K., Huang, J., Zhou, T., Sheng, Z., Kang, B.H., Ishida, E., Griesman, T., Stuccio, S., Bolkhovitinov, L., Wohlbold, T.J., Chromikova, V., Cagigi, A., Leung, K., Andrews, S., Cheung, C.S.F., Pullano, A.A., Plyler, J., Soto, C., Zhang, B., Yang, Y., Joyce, M.G., Tsybovsky, Y., Wheatley, A., Narpala, S.R., Guo, Y., Darko, S., Bailer, R.T., Poole, A., Liang, C.J., Smith, J., Alexander, J., Gurwith, M., Migueles, S.A., Koup, R.A., Golding, H., Khurana, S., McDermott, A.B., Shapiro, L., Krammer, F., Kwong, P.D., Connors, M.(2019) Sci Immunol 4
- PubMed: 31004012
- DOI: https://doi.org/10.1126/sciimmunol.aau2710
- Primary Citation of Related Structures:
6NZ7 - PubMed Abstract:
Induction of an antibody response capable of recognizing highly diverse strains is a major obstacle to the development of vaccines for viruses such as HIV and influenza. Here, we report the dynamics of B cell expansion and evolution at the single-cell level after vaccination with a replication-competent adenovirus type 4 recombinant virus expressing influenza H5 hemagglutinin. Fluorescent H1 or H5 probes were used to quantitate and isolate peripheral blood B cells and their antigen receptors. We observed increases in H5-specific antibody somatic hypermutation and potency for several months beyond the period of active viral replication that was not detectable at the serum level. Individual broad and potent antibodies could be isolated, including one stem-specific antibody that is part of a new multidonor class. These results demonstrate prolonged evolution of the B cell response for months after vaccination and should be considered in efforts to evaluate or boost vaccine-induced immunity.
Organizational Affiliation:
HIV-Specific Immunity Section of the Laboratory of Immunoregulation, National Institutes of Health (NIH), Bethesda, MD 20892, USA.