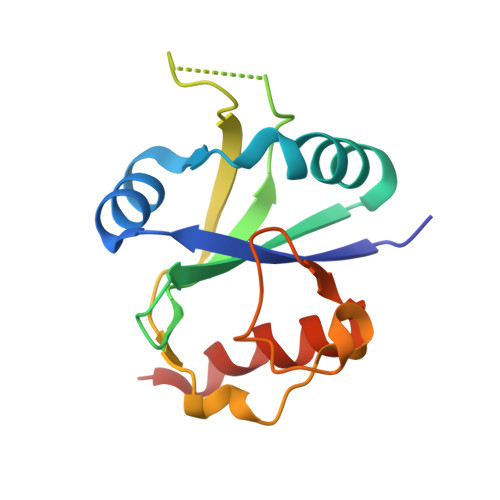
Crystal structure of human YTHDC2 YTH domain.
Ma, C., Liao, S., Zhu, Z.(2019) Biochem Biophys Res Commun 518: 678-684
- PubMed: 31472957
- DOI: https://doi.org/10.1016/j.bbrc.2019.08.107
- Primary Citation of Related Structures:
6K6U - PubMed Abstract:
N 6 -methyladenosine (m 6 A) "readers" play an important role in mRNA functions and metabolism. YTHDC2, as one of the m 6 A readers, controls fertileness through decreasing associated mRNA abundance and enhancing the translation efficiency of related mRNA via binding the targeted m 6 A RNA. However, how YTH domain of YTHDC2 recognize m 6 A RNA is still unknown. In this study, we determined the crystal structure of human YTHDC2 YTH domain, which adopts similar architecture to other solved YTH domain structures. YTHDC2 contains a conserved m 6 A binding pocket, and similar RNA binding surface shared by YTHDC1.
Organizational Affiliation:
Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, 230027, Hefei, China.