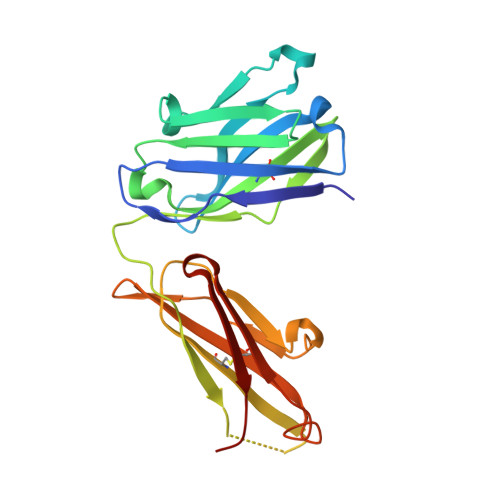
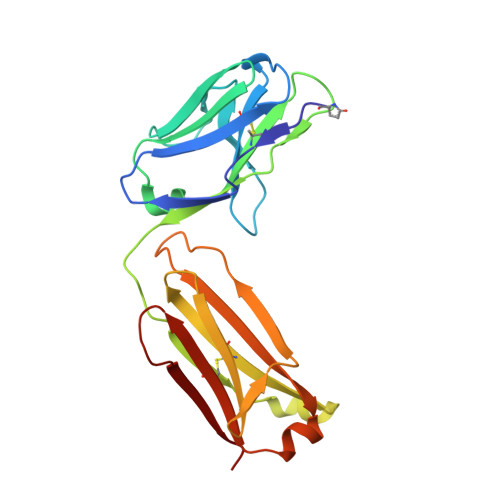
Structural basis of polyethylene glycol recognition by antibody.
Lee, C.C., Su, Y.C., Ko, T.P., Lin, L.L., Yang, C.Y., Chang, S.S., Roffler, S.R., Wang, A.H.(2020) J Biomed Sci 27: 12-12
- PubMed: 31907057
- DOI: https://doi.org/10.1186/s12929-019-0589-7
- Primary Citation of Related Structures:
6JP7, 6JU0, 6JWC - PubMed Abstract:
Polyethylene glycol (PEG) is widely used in industry and medicine. Anti-PEG antibodies have been developed for characterizing PEGylated drugs and other applications. However, the underlying mechanism for specific PEG binding has not been elucidated.
Organizational Affiliation:
Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan. chengung@gate.sinica.edu.tw.