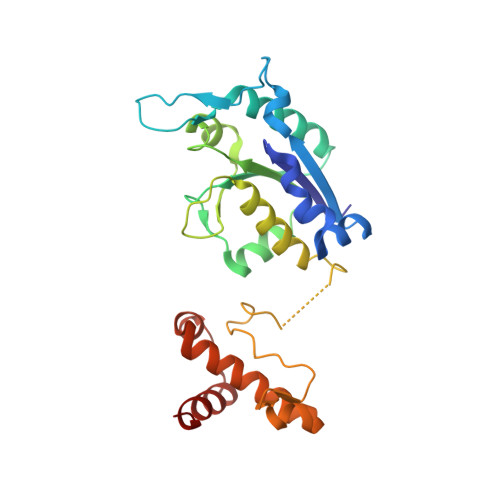
Crystal structure and catalytic mechanism of the essential m1G37 tRNA methyltransferase TrmD fromPseudomonas aeruginosa.
Jaroensuk, J., Wong, Y.H., Zhong, W., Liew, C.W., Maenpuen, S., Sahili, A.E., Atichartpongkul, S., Chionh, Y.H., Nah, Q., Thongdee, N., McBee, M.E., Prestwich, E.G., DeMott, M.S., Chaiyen, P., Mongkolsuk, S., Dedon, P.C., Lescar, J., Fuangthong, M.(2019) RNA 25: 1481-1496
- PubMed: 31399541
- DOI: https://doi.org/10.1261/rna.066746.118
- Primary Citation of Related Structures:
5WYQ, 5WYR, 6JKI - PubMed Abstract:
The tRNA (m 1 G37) methyltransferase TrmD catalyzes m 1 G formation at position 37 in many tRNA isoacceptors and is essential in most bacteria, which positions it as a target for antibiotic development. In spite of its crucial role, little is known about TrmD in Pseudomonas aeruginosa ( Pa TrmD), an important human pathogen. Here we present detailed structural, substrate, and kinetic properties of Pa TrmD. The mass spectrometric analysis confirmed the G36G37-containing tRNAs Leu(GAG), Leu(CAG), Leu(UAG), Pro(GGG), Pro(UGG), Pro(CGG), and His(GUG) as Pa TrmD substrates. Analysis of steady-state kinetics with S -adenosyl-l-methionine (SAM) and tRNA Leu(GAG) showed that Pa TrmD catalyzes the two-substrate reaction by way of a ternary complex, while isothermal titration calorimetry revealed that SAM and tRNA Leu(GAG) bind to Pa TrmD independently, each with a dissociation constant of 14 ± 3 µM. Inhibition by the SAM analog sinefungin was competitive with respect to SAM ( K i = 0.41 ± 0.07 µM) and uncompetitive for tRNA ( K i = 6.4 ± 0.8 µM). A set of crystal structures of the homodimeric Pa TrmD protein bound to SAM and sinefungin provide the molecular basis for enzyme competitive inhibition and identify the location of the bound divalent ion. These results provide insights into Pa TrmD as a potential target for the development of antibiotics.
Organizational Affiliation:
Applied Biological Sciences Program, Chulabhorn Graduate Institute, Chulabhorn Royal Academy, Bangkok 10210, Thailand.