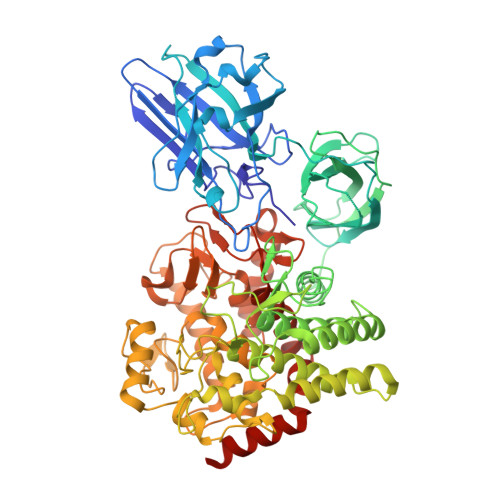
Characterization and Structural Analysis of a Novel exo-Type Enzyme Acting on beta-1,2-Glucooligosaccharides from Parabacteroides distasonis
Shimizu, H., Nakajima, M., Miyanaga, A., Takahashi, Y., Tanaka, N., Kobayashi, K., Sugimoto, N., Nakai, H., Taguchi, H.(2018) Biochemistry 57: 3849-3860
- PubMed: 29763309
- DOI: https://doi.org/10.1021/acs.biochem.8b00385
- Primary Citation of Related Structures:
5Z06 - PubMed Abstract:
β-1,2-Glucan is a polysaccharide produced mainly by some Gram-negative bacteria as a symbiosis and infectious factor. We recently identified endo-β-1,2-glucanase from Chitinophaga pinensis ( CpSGL) as an enzyme comprising a new family. Here, we report the characteristics and crystal structure of a CpSGL homologue from Parabacteroides distasonis, an intestinal bacterium (BDI_3064 protein), which exhibits distinctive properties of known β-1,2-glucan-degrading enzymes. BDI_3064 hydrolyzed linear β-1,2-glucan and β-1,2-glucooligosaccharides with degrees of polymerization (DPs) of ≥4 to produce sophorose specifically but did not hydrolyze cyclic β-1,2-glucan. This result indicates that BDI_3064 is a new exo-type enzyme. BDI_3064 also produced sophorose from β-1,2-glucooligosaccharide analogues that have a modified reducing end, indicating that BDI_3064 acts on its substrates from the nonreducing end. The crystal structure showed that BDI_3064 possesses additional N-terminal domains 1 and 2, unlike CpSGL. Superimposition of BDI_3064 and CpSGL complexed with ligands showed that R93 in domain 1 overlapped subsite -3 in CpSGL. Docking analysis involving a β-1,2-glucooligosaccharide with DP4 showed that R93 completely blocks the nonreducing end of the docked β-1,2-glucooligosaccharide. This indicates that BDI_3064 employs a distinct mechanism of recognition at the nonreducing end of substrates to act as an exo-type enzyme. Thus, we propose 2-β-d-glucooligosaccharide sophorohydrolase (nonreducing end) as a systematic name for BDI_3064.
Organizational Affiliation:
Department of Applied Biological Science, Faculty of Science and Technology , Tokyo University of Science , 2641 Yamazaki , Noda , Chiba 278-8510 , Japan.