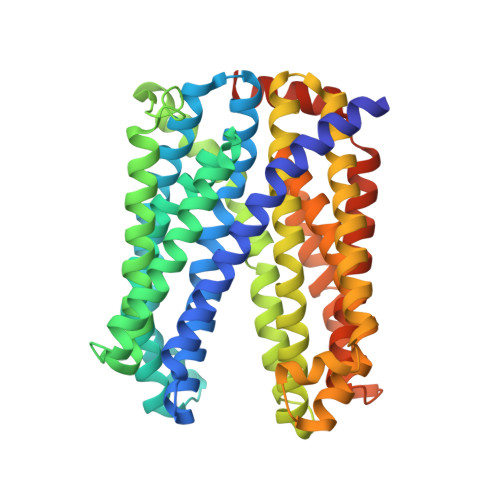
Crystal Structure of a Plant Multidrug and Toxic Compound Extrusion Family Protein
Tanaka, Y., Iwaki, S., Tsukazaki, T.(2017) Structure 25: 1455-1460.e2
- PubMed: 28877507
- DOI: https://doi.org/10.1016/j.str.2017.07.009
- Primary Citation of Related Structures:
5XJJ, 5YCK - PubMed Abstract:
The multidrug and toxic compound extrusion (MATE) family of proteins consists of transporters responsible for multidrug resistance in prokaryotes. In plants, a number of MATE proteins were identified by recent genomic and functional studies, which imply that the proteins have substrate-specific transport functions instead of multidrug extrusion. The three-dimensional structure of eukaryotic MATE proteins, including those of plants, has not been reported, preventing a better understanding of the molecular mechanism of these proteins. Here, we describe the crystal structure of a MATE protein from the plant Camelina sativa at 2.9 Å resolution. Two sets of six transmembrane α helices, assembled pseudo-symmetrically, possess a negatively charged internal pocket with an outward-facing shape. The crystal structure provides insight into the diversity of plant MATE proteins and their substrate recognition and transport through the membrane.
Organizational Affiliation:
Graduate School of Biological Sciences, Nara Institute of Science and Technology, 8916-5 Takayama-cho, Ikoma, Nara 630-0192, Japan.