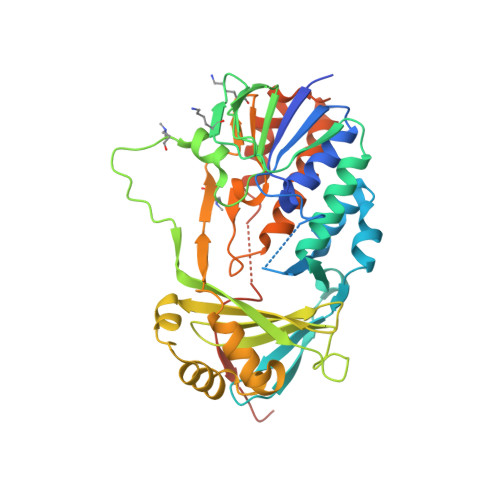
Structural Basis for Inhibitor-Induced Hydrogen Peroxide Production by Kynurenine 3-Monooxygenase
Kim, H.T., Na, B.K., Chung, J., Kim, S., Kwon, S.K., Cha, H., Son, J., Cho, J.M., Hwang, K.Y.(2018) Cell Chem Biol 25: 426-438.e4
- PubMed: 29429898
- DOI: https://doi.org/10.1016/j.chembiol.2018.01.008
- Primary Citation of Related Structures:
5X68, 5X6P, 5X6Q, 5X6R - PubMed Abstract:
Kynurenine 3-monooxygenase (KMO) inhibitors have been developed for the treatment of neurodegenerative disorders. The mechanisms of flavin reduction and hydrogen peroxide production by KMO inhibitors are unknown. Herein, we report the structure of human KMO and crystal structures of Saccharomyces cerevisiae (sc) and Pseudomonas fluorescens (pf) KMO with Ro 61-8048. Proton transfer in the hydrogen bond network triggers flavin reduction in p-hydroxybenzoate hydroxylase, but the mechanism triggering flavin reduction in KMO is different. Conformational changes via π-π interactions between the loop above the flavin and substrate or non-substrate effectors lead to disorder of the C-terminal α helix in scKMO and shifts of domain III in pfKMO, stimulating flavin reduction. Interestingly, Ro 61-8048 has two different binding modes. It acts as a competitive inhibitor in scKMO and as a non-substrate effector in pfKMO. These findings provide understanding of the catalytic cycle of KMO and insight for structure-based drug design of KMO inhibitors.
Organizational Affiliation:
Crystalgenomics, Inc., 5F, Tower A, Korea Bio Park 700, Daewangpangyo-ro, Bundang-gu, Seongnam-si, Gyeonggi-do 13524, Korea; Division of Biotechnology, College of Life Sciences and Biotechnology, Korea University, 145 Anam-ro, Seongbuk-gu, Seoul 02841, Korea.