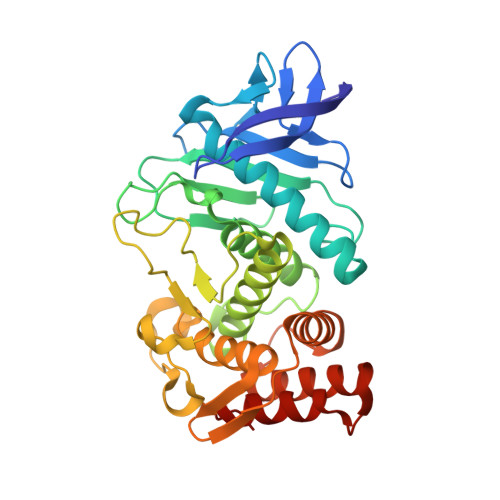
Protein-ligand complex structure from serial femtosecond crystallography using soaked thermolysin microcrystals and comparison with structures from synchrotron radiation
Naitow, H., Matsuura, Y., Tono, K., Joti, Y., Kameshima, T., Hatsui, T., Yabashi, M., Tanaka, R., Tanaka, T., Sugahara, M., Kobayashi, J., Nango, E., Iwata, S., Kunishima, N.(2017) Acta Crystallogr D Struct Biol 73: 702-709
- PubMed: 28777085
- DOI: https://doi.org/10.1107/S2059798317008919
- Primary Citation of Related Structures:
5WR2, 5WR3, 5WR4, 5WR5, 5WR6 - PubMed Abstract:
Serial femtosecond crystallography (SFX) with an X-ray free-electron laser is used for the structural determination of proteins from a large number of microcrystals at room temperature. To examine the feasibility of pharmaceutical applications of SFX, a ligand-soaking experiment using thermolysin microcrystals has been performed using SFX. The results were compared with those from a conventional experiment with synchrotron radiation (SR) at 100 K. A protein-ligand complex structure was successfully obtained from an SFX experiment using microcrystals soaked with a small-molecule ligand; both oil-based and water-based crystal carriers gave essentially the same results. In a comparison of the SFX and SR structures, clear differences were observed in the unit-cell parameters, in the alternate conformation of side chains, in the degree of water coordination and in the ligand-binding mode.
Organizational Affiliation:
Bio-Specimen Platform Group, RIKEN SPring-8 Center, Kouto, Sayo-cho, Sayo-gun, Hyogo 679-5148, Japan.