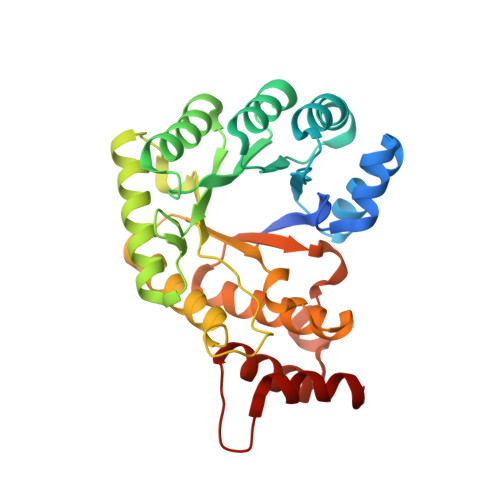
The folate-binding module of Thermus thermophilus cobalamin-dependent methionine synthase displays a distinct variation of the classical TIM barrel: a TIM barrel with a `twist'.
Yamada, K., Koutmos, M.(2018) Acta Crystallogr D Struct Biol 74: 41-51
- PubMed: 29372898
- DOI: https://doi.org/10.1107/S2059798317018290
- Primary Citation of Related Structures:
5VON, 5VOO, 5VOP - PubMed Abstract:
Methyl transfer between methyltetrahydrofolate and corrinoid molecules is a key reaction in biology that is catalyzed by a number of enzymes in many prokaryotic and eukaryotic organisms. One classic example of such an enzyme is cobalamin-dependent methionine synthase (MS). MS is a large modular protein that utilizes an S N 2-type mechanism to catalyze the chemically challenging methyl transfer from the tertiary amine (N5) of methyltetrahydrofolate to homocysteine in order to form methionine. Despite over half a century of study, many questions remain about how folate-dependent methyltransferases, and MS in particular, function. Here, the structure of the folate-binding (Fol) domain of MS from Thermus thermophilus is reported in the presence and absence of methyltetrahydrofolate. It is found that the methyltetrahydrofolate-binding environment is similar to those of previously described methyltransferases, highlighting the conserved role of this domain in binding, and perhaps activating, the methyltetrahydrofolate substrate. These structural studies further reveal a new distinct and uncharacterized topology in the C-terminal region of MS Fol domains. Furthermore, it is found that in contrast to the canonical TIM-barrel β 8 α 8 fold found in all other folate-binding domains, MS Fol domains exhibit a unique β 8 α 7 fold. It is posited that these structural differences are important for MS function.
Organizational Affiliation:
Department of Chemistry, University of Michigan, Ann Arbor, Michigan, USA.