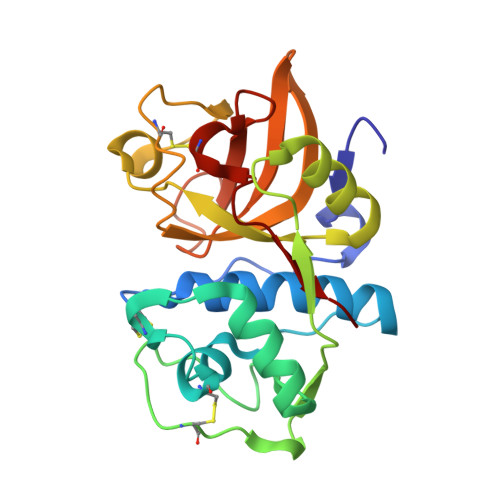
Identification of mouse cathepsin K structural elements that regulate the potency of odanacatib.
Law, S., Andrault, P.M., Aguda, A.H., Nguyen, N.T., Kruglyak, N., Brayer, G.D., Bromme, D.(2017) Biochem J 474: 851-864
- PubMed: 28049758
- DOI: https://doi.org/10.1042/BCJ20160985
- Primary Citation of Related Structures:
5TDI, 5TUN - PubMed Abstract:
Cathepsin K (CatK) is the predominant mammalian bone-degrading protease and thus an ideal target for antiosteoporotic drug development. Rodent models of osteoporosis are preferred due to their close reflection of the human disease and their ease of handling, genetic manipulation and economic affordability. However, large differences in the potency of CatK inhibitors for the mouse/rat vs. the human protease orthologs have made it impossible to use rodent models. This is even more of a problem considering that the most advanced CatK inhibitors, including odanacatib (ODN) and balicatib, failed in human clinical trials due to side effects and rodent models are not available to investigate the mechanism of these failures. Here, we elucidated the structural elements of the potency differences between mouse and human CatK (hCatK) using ODN. We determined and compared the structures of inhibitor-free mouse CatK (mCatK), hCatK and ODN bound to hCatK. Two structural differences were identified and investigated by mutational analysis. Humanizing subsite 2 in mCatK led to a 5-fold improvement of ODN binding, whereas the replacement of Tyr61 in mCatK with Asp resulted in an hCatK with comparable ODN potency. Combining both sites further improved the inhibition of the mCatK variant. Similar results were obtained for balicatib. These findings will allow the generation of transgenic CatK mice that will facilitate the evaluation of CatK inhibitor adverse effects and to explore routes to avoid them.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Faculty of Medicine, University of British Columbia, Vancouver, BC, Canada V6T 1Z3.