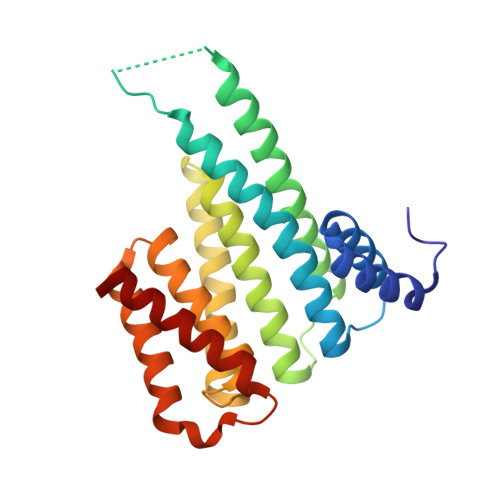
Molecular tweezers modulate 14-3-3 protein-protein interactions.
Bier, D., Rose, R., Bravo-Rodriguez, K., Bartel, M., Ramirez-Anguita, J.M., Dutt, S., Wilch, C., Klarner, F.G., Sanchez-Garcia, E., Schrader, T., Ottmann, C.(2013) Nat Chem 5: 234-239
- PubMed: 23422566
- DOI: https://doi.org/10.1038/nchem.1570
- Primary Citation of Related Structures:
5OEG, 5OEH - PubMed Abstract:
Supramolecular chemistry has recently emerged as a promising way to modulate protein functions, but devising molecules that will interact with a protein in the desired manner is difficult as many competing interactions exist in a biological environment (with solvents, salts or different sites for the target biomolecule). We now show that lysine-specific molecular tweezers bind to a 14-3-3 adapter protein and modulate its interaction with partner proteins. The tweezers inhibit binding between the 14-3-3 protein and two partner proteins--a phosphorylated (C-Raf) protein and an unphosphorylated one (ExoS)--in a concentration-dependent manner. Protein crystallography shows that this effect arises from the binding of the tweezers to a single surface-exposed lysine (Lys214) of the 14-3-3 protein in the proximity of its central channel, which normally binds the partner proteins. A combination of structural analysis and computer simulations provides rules for the tweezers' binding preferences, thus allowing us to predict their influence on this type of protein-protein interactions.
Organizational Affiliation:
Chemical Genomics Centre of the Max-Planck-Society, Otto-Hahn-Strasse 15, 44227 Dortmund, Germany.