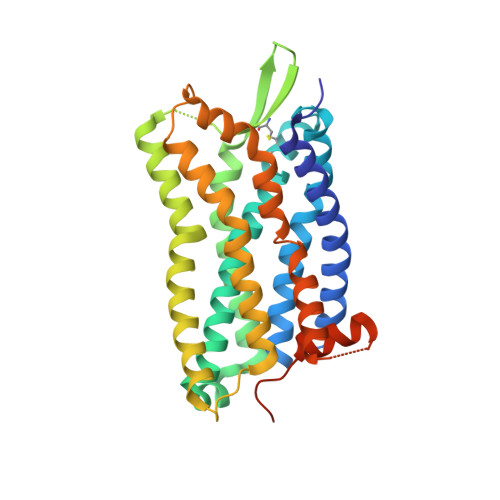
Structure of the complement C5a receptor bound to the extra-helical antagonist NDT9513727.
Robertson, N., Rappas, M., Dore, A.S., Brown, J., Bottegoni, G., Koglin, M., Cansfield, J., Jazayeri, A., Cooke, R.M., Marshall, F.H.(2018) Nature 553: 111-114
- PubMed: 29300009
- DOI: https://doi.org/10.1038/nature25025
- Primary Citation of Related Structures:
5O9H - PubMed Abstract:
The complement system is a crucial component of the host response to infection and tissue damage. Activation of the complement cascade generates anaphylatoxins including C5a and C3a. C5a exerts a pro-inflammatory effect via the G-protein-coupled receptor C5a anaphylatoxin chemotactic receptor 1 (C5aR1, also known as CD88) that is expressed on cells of myeloid origin. Inhibitors of the complement system have long been of interest as potential drugs for the treatment of diseases such as sepsis, rheumatoid arthritis, Crohn's disease and ischaemia-reperfusion injuries. More recently, a role of C5a in neurodegenerative conditions such as Alzheimer's disease has been identified. Peptide antagonists based on the C5a ligand have progressed to phase 2 trials in psoriasis and rheumatoid arthritis; however, these compounds exhibited problems with off-target activity, production costs, potential immunogenicity and poor oral bioavailability. Several small-molecule competitive antagonists for C5aR1, such as W-54011 and NDT9513727, have been identified by C5a radioligand-binding assays. NDT9513727 is a non-peptide inverse agonist of C5aR1, and is highly selective for the primate and gerbil receptors over those of other species. Here, to study the mechanism of action of C5a antagonists, we determine the structure of a thermostabilized C5aR1 (known as C5aR1 StaR) in complex with NDT9513727. We found that the small molecule bound between transmembrane helices 3, 4 and 5, outside the helical bundle. One key interaction between the small molecule and residue Trp213 5.49 seems to determine the species selectivity of the compound. The structure demonstrates that NDT9513727 exerts its inverse-agonist activity through an extra-helical mode of action.
Organizational Affiliation:
Heptares Therapeutics Ltd, BioPark, Broadwater Road, Welwyn Garden City, Hertfordshire AL7 3AX, UK.