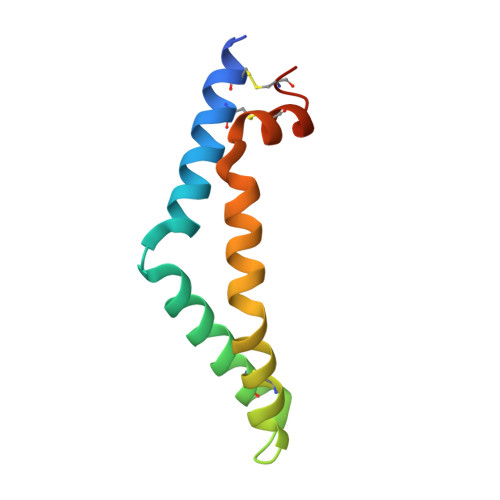
The mechanism of glycosphingolipid degradation revealed by a GALC-SapA complex structure.
Hill, C.H., Cook, G.M., Spratley, S.J., Fawke, S., Graham, S.C., Deane, J.E.(2018) Nat Commun 9: 151-151
- PubMed: 29323104
- DOI: https://doi.org/10.1038/s41467-017-02361-y
- Primary Citation of Related Structures:
5NXB - PubMed Abstract:
Sphingolipids are essential components of cellular membranes and defects in their synthesis or degradation cause severe human diseases. The efficient degradation of sphingolipids in the lysosome requires lipid-binding saposin proteins and hydrolytic enzymes. The glycosphingolipid galactocerebroside is the primary lipid component of the myelin sheath and is degraded by the hydrolase β-galactocerebrosidase (GALC). This enzyme requires the saposin SapA for lipid processing and defects in either of these proteins causes a severe neurodegenerative disorder, Krabbe disease. Here we present the structure of a glycosphingolipid-processing complex, revealing how SapA and GALC form a heterotetramer with an open channel connecting the enzyme active site to the SapA hydrophobic cavity. This structure defines how a soluble hydrolase can cleave the polar glycosyl headgroups of these essential lipids from their hydrophobic ceramide tails. Furthermore, the molecular details of this interaction provide an illustration for how specificity of saposin binding to hydrolases is encoded.
Organizational Affiliation:
Cambridge Institute for Medical Research, Department of Pathology, University of Cambridge, Cambridge Biomedical Campus, Hills Road, Cambridge, CB2 0XY, UK.