New active leads for tuberculosis booster drugs by structure-based drug discovery.
Tatum, N.J., Liebeschuetz, J.W., Cole, J.C., Frita, R., Herledan, A., Baulard, A.R., Willand, N., Pohl, E.(2017) Org Biomol Chem 15: 10245-10255
- PubMed: 29182187
- DOI: https://doi.org/10.1039/c7ob00910k
- Primary Citation of Related Structures:
5NIM, 5NIO, 5NIZ, 5NJ0 - PubMed Abstract:
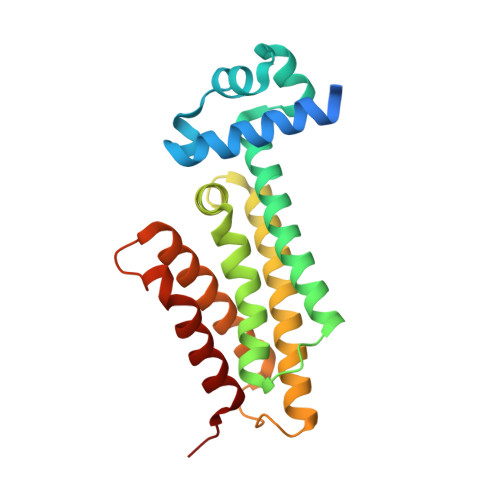
The transcriptional repressor EthR from Mycobacterium tuberculosis, a member of the TetR family of prokaryotic homo-dimeric transcription factors, controls the expression of the mycobacterial mono-oxygenase EthA. EthA is responsible for the bio-activation of the second-line tuberculosis pro-drug ethionamide, and consequently EthR inhibitors boost drug efficacy. Here, we present a comprehensive in silico structure-based screening protocol that led to the identification of a number of novel scaffolds of EthR inhibitors in subsequent biophysical screening by thermal shift assay. Growth inhibition assays demonstrated that five of the twenty biophysical hits were capable of boosting ethionamide activity in vitro, with the best novel scaffold displaying an EC 50 of 34 μM. In addition, the co-crystal structures of EthR with four new ligands at resolution ranging from 2.1 to 1.4 Å confirm the binding and inactivation mode, and will enable future lead development.
Organizational Affiliation:
Department of Chemistry, Durham University, South Road, Durham DH1 3LE, UK. ehmke.pohl@durham.ac.uk.