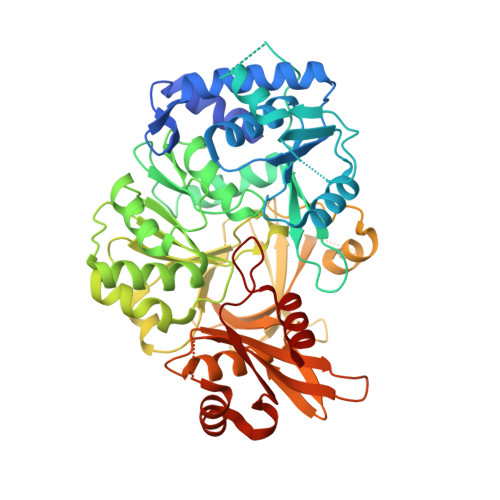
Structure of the adenylation domain Thr1 involved in the biosynthesis of 4-chlorothreonine in Streptomyces sp. OH-5093-protein flexibility and molecular bases of substrate specificity.
Scaglione, A., Fullone, M.R., Montemiglio, L.C., Parisi, G., Zamparelli, C., Vallone, B., Savino, C., Grgurina, I.(2017) FEBS J 284: 2981-2999
- PubMed: 28704585
- DOI: https://doi.org/10.1111/febs.14163
- Primary Citation of Related Structures:
5N9W, 5N9X - PubMed Abstract:
We determined the crystal structure of Thr1, the self-standing adenylation domain involved in the nonribosomal-like biosynthesis of free 4-chlorothreonine in Streptomyces sp. OH-5093. Thr1 shows two monomers in the crystallographic asymmetric unit with different relative orientations of the C- and N-terminal subdomains both in the presence of substrates and in the unliganded form. Cocrystallization with substrates, adenosine 5'-triphosphate and l-threonine, yielded one monomer containing the two substrates and the other in complex with l-threonine adenylate, locked in a postadenylation state. Steady-state kinetics showed that Thr1 activates l-Thr and its stereoisomers, as well as d-Ala, l- and d-Ser, albeit with lower efficiency. Modeling of these substrates in the active site highlighted the molecular bases of substrate discrimination. This work provides the first crystal structure of a threonine-activating adenylation enzyme, a contribution to the studies on conformational rearrangement in adenylation domains and on substrate recognition in nonribosomal biosynthesis.
Organizational Affiliation:
Department of Biochemical Sciences "A. Rossi Fanelli", Istituto Pasteur-Fondazione Cenci Bolognetti, Sapienza University of Rome, Italy.