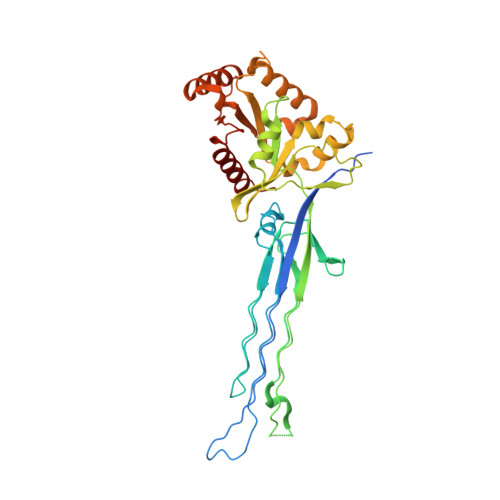
Structural and biochemical analysis of Escherichia coli ObgE, a central regulator of bacterial persistence.
Gkekas, S., Singh, R.K., Shkumatov, A.V., Messens, J., Fauvart, M., Verstraeten, N., Michiels, J., Versees, W.(2017) J Biol Chem 292: 5871-5883
- PubMed: 28223358
- DOI: https://doi.org/10.1074/jbc.M116.761809
- Primary Citation of Related Structures:
5M04 - PubMed Abstract:
The Obg protein family belongs to the TRAFAC (translation factor) class of P-loop GTPases and is conserved from bacteria to eukaryotes. Essential roles in many different cellular processes have been suggested for the Obg protein from Escherichia coli (ObgE), and we recently showed that it is a central regulator of bacterial persistence. Here, we report the first crystal structure of ObgE at 1.85-Å resolution in the GDP-bound state, showing the characteristic N-terminal domain and a central G domain that are common to all Obg proteins. ObgE also contains an intrinsically disordered C-terminal domain, and we show here that this domain specifically contributed to GTP binding, whereas it did not influence GDP binding or GTP hydrolysis. Biophysical analysis, using small angle X-ray scattering and multi-angle light scattering experiments, revealed that ObgE is a monomer in solution, regardless of the bound nucleotide. In contrast to recent suggestions, our biochemical analyses further indicate that ObgE is neither activated by K + ions nor by homodimerization. However, the ObgE GTPase activity was stimulated upon binding to the ribosome, confirming the ribosome-dependent GTPase activity of the Obg family. Combined, our data represent an important step toward further unraveling the detailed molecular mechanism of ObgE, which might pave the way to further studies into how this GTPase regulates bacterial physiology, including persistence.
Organizational Affiliation:
From the Structural Biology Brussels, Vrije Universiteit Brussel, 1050 Brussels.