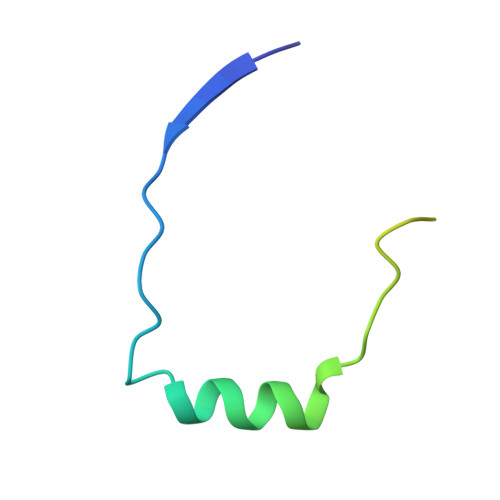
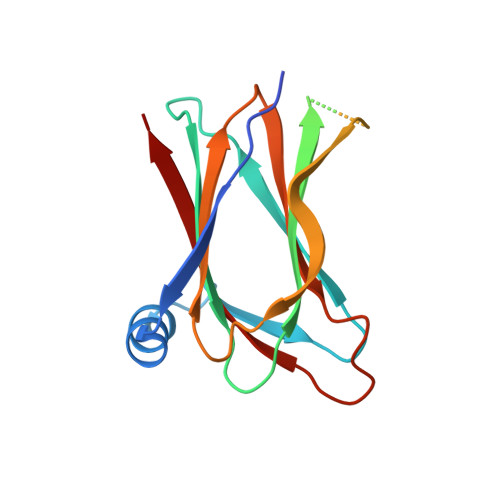
Kobuviral Non-structural 3A Proteins Act as Molecular Harnesses to Hijack the Host ACBD3 Protein.
Klima, M., Chalupska, D., Rozycki, B., Humpolickova, J., Rezabkova, L., Silhan, J., Baumlova, A., Dubankova, A., Boura, E.(2017) Structure 25: 219-230
- PubMed: 28065508
- DOI: https://doi.org/10.1016/j.str.2016.11.021
- Primary Citation of Related Structures:
5LZ1, 5LZ3, 5LZ6 - PubMed Abstract:
Picornaviruses are small positive-sense single-stranded RNA viruses that include many important human pathogens. Within the host cell, they replicate at specific replication sites called replication organelles. To create this membrane platform, they hijack several host factors including the acyl-CoA-binding domain-containing protein-3 (ACBD3). Here, we present a structural characterization of the molecular complexes formed by the non-structural 3A proteins from two species of the Kobuvirus genus of the Picornaviridae family and the 3A-binding domain of the host ACBD3 protein. Specifically, we present a series of crystal structures as well as a molecular dynamics simulation of the 3A:ACBD3 complex at the membrane, which reveals that the viral 3A proteins act as molecular harnesses to enslave the ACBD3 protein leading to its stabilization at target membranes. Our data provide a structural rationale for understanding how these viral-host protein complexes assemble at the atomic level and identify new potential targets for antiviral therapies.
Organizational Affiliation:
Institute of Organic Chemistry and Biochemistry, Czech Academy of Sciences, 16610 Prague, Czech Republic. Electronic address: klima@uochb.cas.cz.