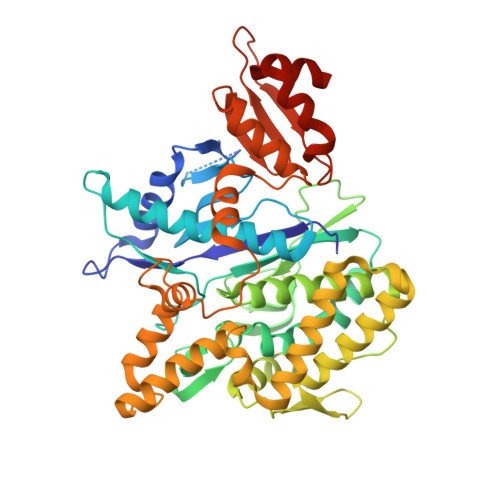
Depupylase Dop Requires Inorganic Phosphate in the Active Site for Catalysis.
Bolten, M., Vahlensieck, C., Lipp, C., Leibundgut, M., Ban, N., Weber-Ban, E.(2017) J Biol Chem 292: 4044-4053
- PubMed: 28119453
- DOI: https://doi.org/10.1074/jbc.M116.755645
- Primary Citation of Related Structures:
5LRT - PubMed Abstract:
Analogous to eukaryotic ubiquitination, proteins in actinobacteria can be post-translationally modified in a process referred to as pupylation, the covalent attachment of prokaryotic ubiquitin-like protein Pup to lysine side chains of the target protein via an isopeptide bond. As in eukaryotes, an opposing activity counteracts the modification by specific cleavage of the isopeptide bond formed with Pup. However, the enzymes involved in pupylation and depupylation have evolved independently of ubiquitination and are related to the family of ATP-binding and hydrolyzing carboxylate-amine ligases of the glutamine synthetase type. Furthermore, the Pup ligase PafA and the depupylase Dop share close structural and sequence homology and have a common evolutionary history despite catalyzing opposing reactions. Here, we investigate the role played by the nucleotide in the active site of the depupylase Dop using a combination of biochemical experiments and X-ray crystallographic studies. We show that, although Dop does not turn over ATP stoichiometrically with substrate, the active site nucleotide species in Dop is ADP and inorganic phosphate rather than ATP, and that non-hydrolyzable analogs of ATP cannot support the enzymatic reaction. This finding suggests that the catalytic mechanism is more similar to the mechanism of the ligase PafA than previously thought and likely involves the transient formation of a phosphorylated Pup-intermediate. Evidence is presented for a mechanism where the inorganic phosphate acts as the nucleophilic species in amide bond cleavage and implications for Dop function are discussed.
Organizational Affiliation:
From the ETH Zurich, Institute of Molecular Biology & Biophysics, 8093 Zurich, Switzerland.