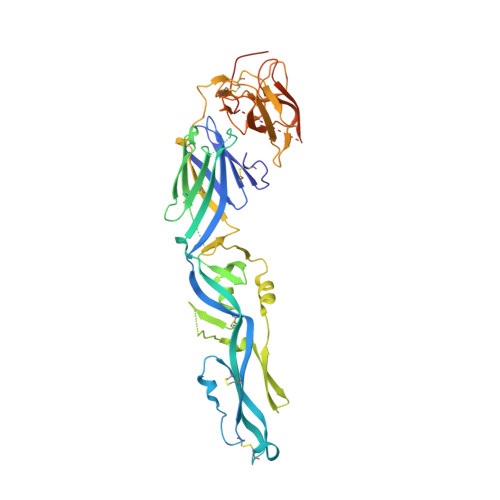
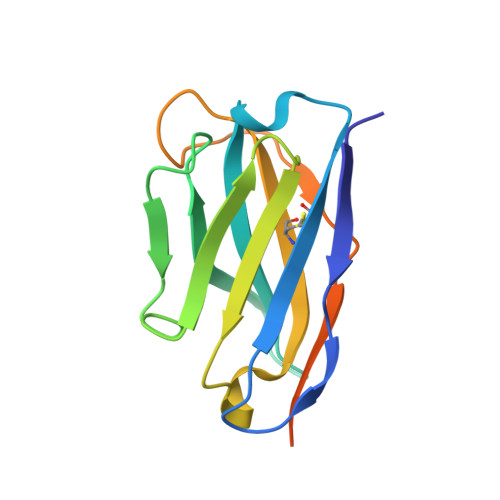
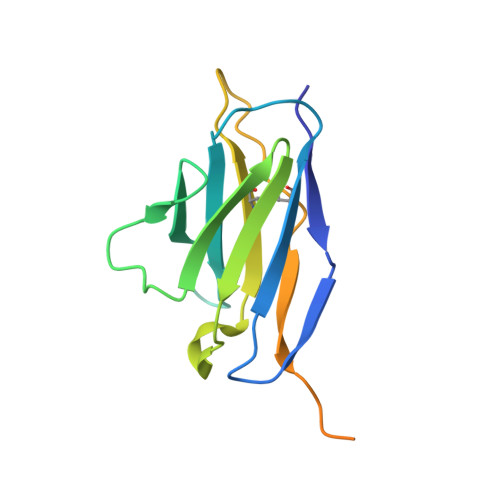
Structural basis of potent Zika-dengue virus antibody cross-neutralization.
Barba-Spaeth, G., Dejnirattisai, W., Rouvinski, A., Vaney, M.C., Medits, I., Sharma, A., Simon-Loriere, E., Sakuntabhai, A., Cao-Lormeau, V.M., Haouz, A., England, P., Stiasny, K., Mongkolsapaya, J., Heinz, F.X., Screaton, G.R., Rey, F.A.(2016) Nature 536: 48-53
- PubMed: 27338953
- DOI: https://doi.org/10.1038/nature18938
- Primary Citation of Related Structures:
5LBS, 5LCV - PubMed Abstract:
Zika virus is a member of the Flavivirus genus that had not been associated with severe disease in humans until the recent outbreaks, when it was linked to microcephaly in newborns in Brazil and to Guillain-Barré syndrome in adults in French Polynesia. Zika virus is related to dengue virus, and here we report that a subset of antibodies targeting a conformational epitope isolated from patients with dengue virus also potently neutralize Zika virus. The crystal structure of two of these antibodies in complex with the envelope protein of Zika virus reveals the details of a conserved epitope, which is also the site of interaction of the envelope protein dimer with the precursor membrane (prM) protein during virus maturation. Comparison of the Zika and dengue virus immunocomplexes provides a lead for rational, epitope-focused design of a universal vaccine capable of eliciting potent cross-neutralizing antibodies to protect simultaneously against both Zika and dengue virus infections.