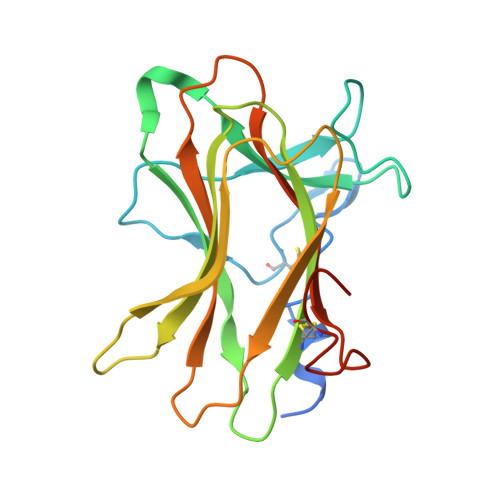
Crystal Structure of the Neuropilin-1 MAM Domain: Completing the Neuropilin-1 Ectodomain Picture.
Yelland, T., Djordjevic, S.(2016) Structure 24: 2008-2015
- PubMed: 27720589
- DOI: https://doi.org/10.1016/j.str.2016.08.017
- Primary Citation of Related Structures:
5L73 - PubMed Abstract:
Neuropilins (NRPs) are single-pass transmembrane receptors involved in several signaling pathways that regulate key physiological processes such as vascular morphogenesis and axon guidance. The MAM domain of NRP, which has previously been implicated in receptor multimerization, was the only portion of the ectopic domain of the NRPs for which the structure, until now, has been elusive. Using site-directed mutagenesis in the linker region preceding the MAM domain we generated a protein construct amenable to crystallization. Here we present the crystal structure of the MAM domain of human NRP1 at 2.24 Å resolution. The protein exhibits a jellyroll topology, with Ca 2+ ions bound at the inter-strand space enhancing the thermostability of the domain. We show that the MAM domain of NRP1 is monomeric in solution and insufficient to drive receptor dimerization, which leads us to propose a different role for this domain in the context of NRP membrane assembly and signaling.
Organizational Affiliation:
The Institute of Structural and Molecular Biology, University College London, Gower Street, London WC1E 6BT, UK.