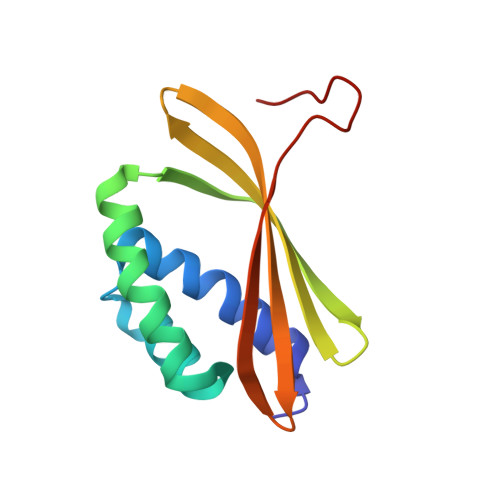
Principles for designing proteins with cavities formed by curved beta sheets.
Marcos, E., Basanta, B., Chidyausiku, T.M., Tang, Y., Oberdorfer, G., Liu, G., Swapna, G.V., Guan, R., Silva, D.A., Dou, J., Pereira, J.H., Xiao, R., Sankaran, B., Zwart, P.H., Montelione, G.T., Baker, D.(2017) Science 355: 201-206
- PubMed: 28082595
- DOI: https://doi.org/10.1126/science.aah7389
- Primary Citation of Related Structures:
5KPE, 5KPH, 5L33, 5TPH, 5TPJ, 5TRV, 5TS4, 5U35 - PubMed Abstract:
Active sites and ligand-binding cavities in native proteins are often formed by curved β sheets, and the ability to control β-sheet curvature would allow design of binding proteins with cavities customized to specific ligands. Toward this end, we investigated the mechanisms controlling β-sheet curvature by studying the geometry of β sheets in naturally occurring protein structures and folding simulations. The principles emerging from this analysis were used to design, de novo, a series of proteins with curved β sheets topped with α helices. Nuclear magnetic resonance and crystal structures of the designs closely match the computational models, showing that β-sheet curvature can be controlled with atomic-level accuracy. Our approach enables the design of proteins with cavities and provides a route to custom design ligand-binding and catalytic sites.
Organizational Affiliation:
Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.