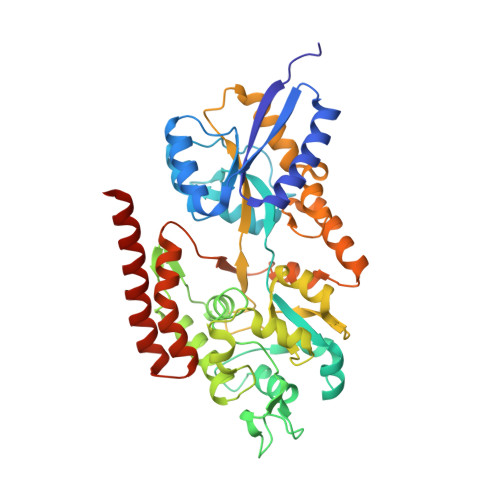
Structural and functional analysis of the solute-binding protein UspC from Mycobacterium tuberculosis that is specific for amino sugars.
Fullam, E., Prokes, I., Futterer, K., Besra, G.S.(2016) Open Biol 6
- PubMed: 27335320
- DOI: https://doi.org/10.1098/rsob.160105
- Primary Citation of Related Structures:
5K2X, 5K2Y - PubMed Abstract:
Mycobacterium tuberculosis (Mtb), the aetiological agent of tuberculosis, has evolved to scavenge nutrients from the confined environment of host macrophages with mycobacterial ATP-binding cassette (ABC) transporters playing a key role in nutrient acquisition. Mtb-UspC (Rv2318) is the solute-binding protein of the essential transporter UspABC, one of four Mtb ABC transporters implicated by homology in sugar acquisition. Herein, we report the structural and functional characterization of Mtb-UspC. The 1.5 Å resolution structure of UspC reveals a two subdomain architecture that forms a highly acidic carbohydrate-substrate binding cleft. This has allowed a distinct preference of Mtb-UspC for amino sugars as determined by thermal shift analysis and solution saturation transfer difference-NMR. Taken together our data support the functional assignment of UspABC as an amino-sugar transporter. Given the limited availability of carbohydrates within the phagosomal environmental niche during Mtb intracellular infection, our studies suggest that UspABC enables Mtb to optimize the use of scarce nutrients during intracellular infection, linking essentiality of this protein to a potential role in recycling components of cell-wall peptidoglycan.
Organizational Affiliation:
School of Life Sciences, University of Warwick, Coventry CV4 7AL, UK School of Biosciences, University of Birmingham, Edgbaston, Birmingham B15 2TT, UK e.fullam@warwick.ac.uk.