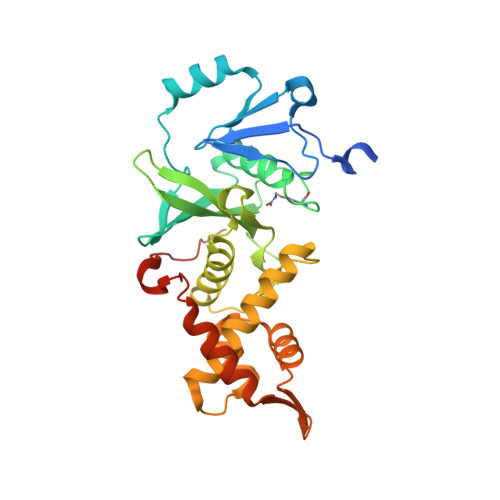
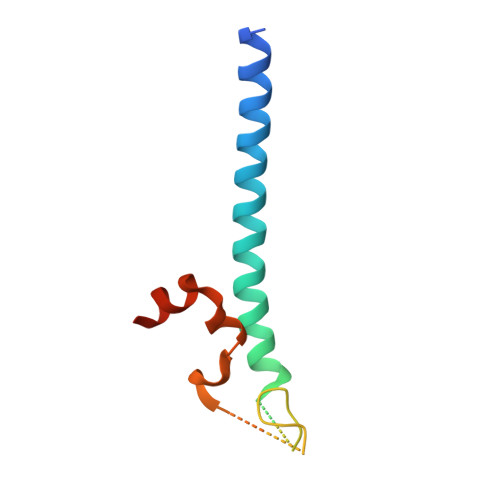
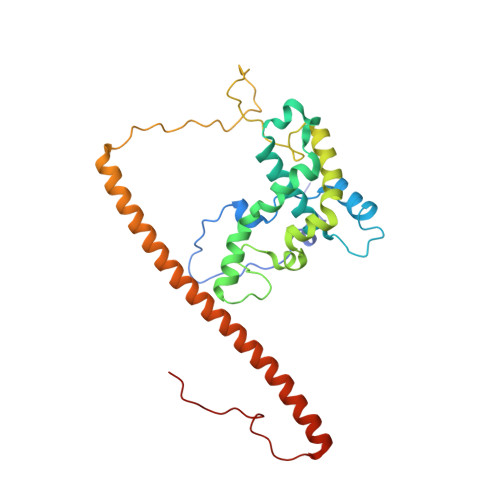
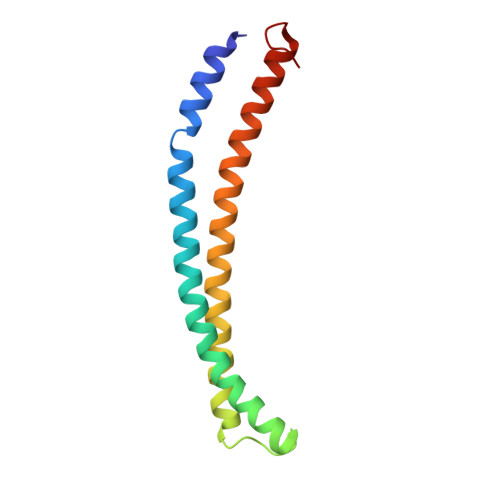
The NuA4 Core Complex Acetylates Nucleosomal Histone H4 through a Double Recognition Mechanism
Xu, P., Li, C., Chen, Z., Jiang, S., Fan, S., Wang, J., Dai, J., Zhu, P., Chen, Z.(2016) Mol Cell 63: 965-975
- PubMed: 27594449
- DOI: https://doi.org/10.1016/j.molcel.2016.07.024
- Primary Citation of Related Structures:
5J9Q, 5J9T, 5J9U, 5J9W - PubMed Abstract:
NuA4 catalyzes the acetylation of nucleosomes at histone H4, which is a well-established epigenetic event, controlling many genomic processes in Saccharomyces cerevisiae. Here we report the crystal structures of the NuA4 core complex and a cryoelectron microscopy structure with the nucleosome. The structures show that the histone-binding pocket of the enzyme is rearranged, suggesting its activation. The enzyme binds the histone tail mainly through the target lysine residue, with a preference for a small residue at the -1 position. The complex engages the nucleosome at the dish face and orients its catalytic pocket close to the H4 tail to achieve selective acetylation. The combined data reveal a space-sequence double recognition mechanism of the histone tails by a modifying enzyme in the context of the nucleosome.
Organizational Affiliation:
MOE Key Laboratory of Protein Science, Tsinghua University, Beijing 100086, China; School of Life Science, Tsinghua University, Beijing 100086, China.