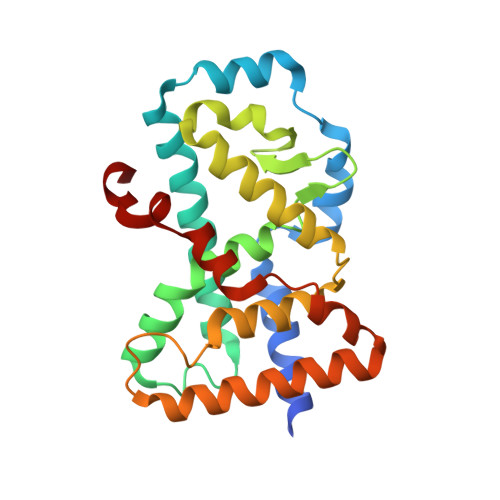
Structural determinant for inducing RORgamma specific inverse agonism triggered by a synthetic benzoxazinone ligand.
Marcotte, D.J., Liu, Y., Little, K., Jones, J.H., Powell, N.A., Wildes, C.P., Silvian, L.F., Chodaparambil, J.V.(2016) BMC Struct Biol 16: 7-7
- PubMed: 27246200
- DOI: https://doi.org/10.1186/s12900-016-0059-3
- Primary Citation of Related Structures:
5IXK, 5IZ0 - PubMed Abstract:
The nuclear hormone receptor RORγ regulates transcriptional genes involved in the production of the pro-inflammatory interleukin IL-17 which has been linked to autoimmune diseases such as rheumatoid arthritis, multiple sclerosis and inflammatory bowel disease. This transcriptional activity of RORγ is modulated through a protein-protein interaction involving the activation function 2 (AF2) helix on the ligand binding domain of RORγ and a conserved LXXLL helix motif on coactivator proteins. Our goal was to develop a RORγ specific inverse agonist that would help down regulate pro-inflammatory gene transcription by disrupting the protein protein interaction with coactivator proteins as a therapeutic agent.
Organizational Affiliation:
Chemical and Molecular Therapeutics, Biogen Inc, 250 Binney Street, Cambridge, MA, 02142, USA. doug.marcotte@biogenidec.com.