Staphylococcus aureus Dihydrofolate Reductase complexed with cyclized alpha-NADPH anomer and 3'-(3-(2,4-diamino-6-ethylpyrimidin-5-yl)prop-2-yn-1-yl)-5'-methoxy-[1,1'-biphenyl]-4-carboxylic acid (UCP1175)
Reeve, S.M., Anderson, A.C.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Dihydrofolate reductase | A [auth X] | 157 | Staphylococcus aureus | Mutation(s): 0 Gene Names: folA EC: 1.5.1.3 | 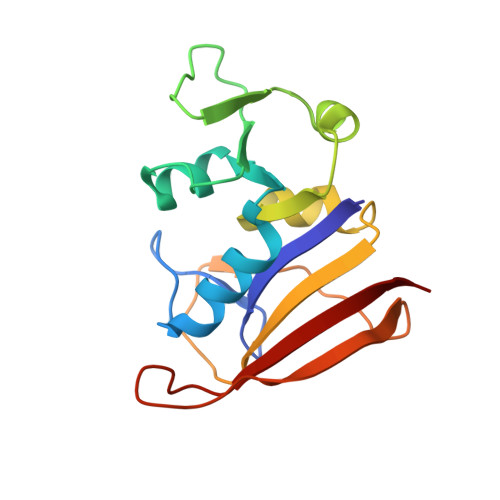 |
UniProt | |||||
Find proteins for P0A017 (Staphylococcus aureus) Explore P0A017 Go to UniProtKB: P0A017 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A017 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| XNP Query on XNP | B [auth X] | Tricyclic NADPH C21 H30 N7 O17 P3 AJKICLDTLPZSPE-MTKBYBFRSA-N |  | ||
| U75 Query on U75 | C [auth X] | 4-[3-[3-[2,4-bis(azanyl)-6-ethyl-pyrimidin-5-yl]prop-2-ynyl]-5-methoxy-phenyl]benzoic acid C23 H22 N4 O3 VLQRVOAJCWFFQG-UHFFFAOYSA-N |  | ||
| GOL Query on GOL | D [auth X] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 79.022 | α = 90 |
| b = 79.022 | β = 90 |
| c = 108.247 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| d*TREK | data reduction |
| StructureStudio | data collection |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | AI111957 |