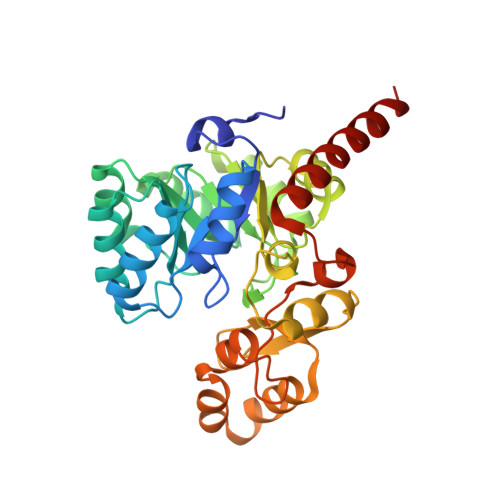
Structural and biochemical characterization of FabK from Thermotoga maritima.
Ha, B.H., Shin, S.C., Moon, J.H., Keum, G., Kim, C.W., Kim, E.E.(2017) Biochem Biophys Res Commun 482: 968-974
- PubMed: 27908729
- DOI: https://doi.org/10.1016/j.bbrc.2016.11.141
- Primary Citation of Related Structures:
5GVH, 5GVJ - PubMed Abstract:
TM0800 from Thermotoga maritima is one of the hypothetical proteins with unknown function. The crystal structure determined at 2.3 Å resolution reveals a two domain structure: the N-terminal domain forming a barrel and the C-terminal forming a lid. One FMN is bound between the two domains with the phosphate making intricate hydrogen bonds with protein and three tightly bound water molecules, and the isoalloxazine ring packed against the side chains of Met22 and Met276. The structure is almost identical to that of FabK (enoyl-acyl carrier protein (ACP) reductase, ENR II), a key enzyme in bacterial type II fatty-acid biosynthesis that catalyzes the final step in each elongation cycle; and the enzymatic activity confirms that TM0800 is an ENR. Enzymatic activity was almost completely abolished when the helices connecting the barrel and the lid were deleted. Also, the Met276Ala and Ser280Ala mutants showed a significant reduction in enzymatic activity. The crystal structure of Met276Ala mutant at 1.9 Å resolution showed an absence of FMN suggesting that FMN plays a role in catalysis, and Met276 is important in positioning FMN. TmFabK exists as a dimer in both solution and crystal. Together this study provides molecular basis for the catalytic activity of FabK.
Organizational Affiliation:
Biomedical Research Institute, Korea Institute of Science and Technology, Seoul, 02792, Republic of Korea.