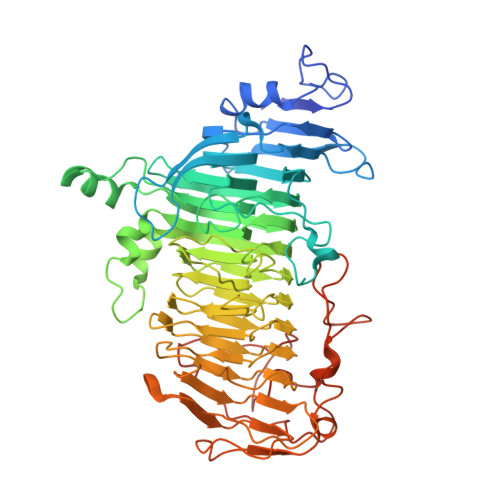
Molecular Insight into Evolution of Symbiosis between Breast-Fed Infants and a Member of the Human Gut Microbiome Bifidobacterium longum
Yamada, C., Gotoh, A., Sakanaka, M., Hattie, M., Stubbs, K.A., Katayama-Ikegami, A., Hirose, J., Kurihara, S., Arakawa, T., Kitaoka, M., Okuda, S., Katayama, T., Fushinobu, S.(2017) Cell Chem Biol 24: 515-524.e5
- PubMed: 28392148
- DOI: https://doi.org/10.1016/j.chembiol.2017.03.012
- Primary Citation of Related Structures:
5GQC, 5GQF, 5GQG - PubMed Abstract:
Breast-fed infants generally have a bifidobacteria-rich microbiota with recent studies indicating that human milk oligosaccharides (HMOs) selectively promote bifidobacterial growth. Bifidobacterium bifidum possesses a glycoside hydrolase family 20 lacto-N-biosidase for liberating lacto-N-biose I from lacto-N-tetraose, an abundant HMO unique to human milk, while Bifidobacterium longum subsp. longum has a non-classified enzyme (LnbX). Here, we determined the crystal structure of the catalytic domain of LnbX and provide evidence for creation of a novel glycoside hydrolase family, GH136. The structure, in combination with inhibition and mutation studies, provides insight into the molecular mechanism and broader substrate specificity of this enzyme. Moreover, through genetic studies, we show that lnbX is indispensable for B. longum growth on lacto-N-tetraose and is a key genetic factor for persistence in the gut of breast-fed infants. Overall, this study reveals possible evolutionary routes for the emergence of symbiosis between humans and bifidobacterial species in the infant gut.
Organizational Affiliation:
Department of Biotechnology, The University of Tokyo, Tokyo 113-8657, Japan; Graduate School of Biostudies, Kyoto University, Kyoto 606-8052, Japan.