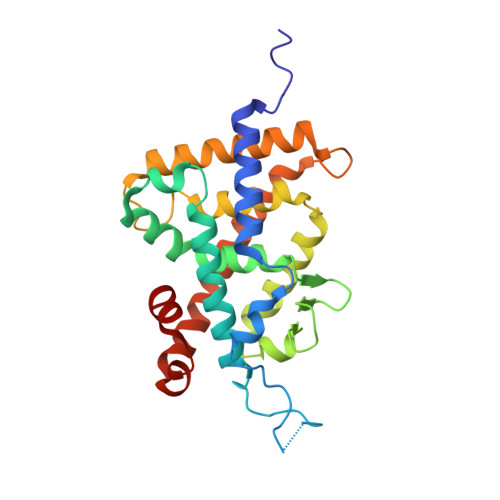
Regulation of the vitamin D receptor by vitamin D lactam derivatives.
Asano, L., Waku, T., Abe, R., Kuwabara, N., Ito, I., Yanagisawa, J., Nagasawa, K., Shimizu, T.(2016) FEBS Lett 590: 3270-3279
- PubMed: 27500498
- DOI: https://doi.org/10.1002/1873-3468.12348
- Primary Citation of Related Structures:
5GIC, 5GID, 5GIE - PubMed Abstract:
The active metabolite of vitamin D3 , 1α,25-dihydroxyvitamin D3 , acts as a ligand for the vitamin D receptor (VDR) and activates VDR-mediated gene expression. Recently, we characterized 1α,25-dihydroxyvitamin D3 -26,23-lactams (DLAMs), which mimic vitamin D3 metabolites, as noncalcemic VDR ligands that barely activate the receptor. In this study, we present structural insights onto the regulation of VDR function by DLAMs. X-ray crystallographic analysis revealed that DLAMs induced a large conformational change in the loop region between helices H6 and H7 in the VDR ligand-binding domain. Our structural analysis suggests that targeting of the loop region may be a new mode of VDR regulation.
Organizational Affiliation:
Institute for Chemical Research, Kyoto University, Uji, Kyoto, Japan.