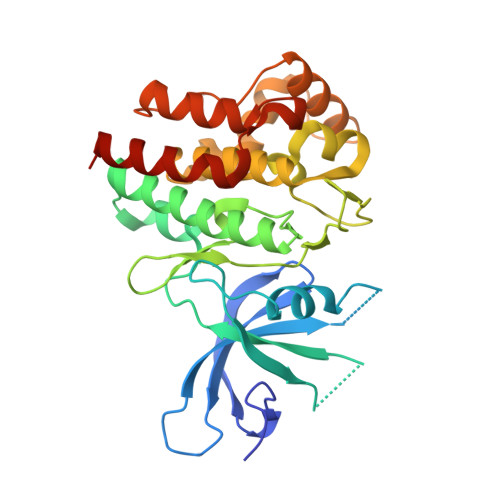
Structure-Based Design and Synthesis of 3-Amino-1,5-dihydro-4H-pyrazolopyridin-4-one Derivatives as Tyrosine Kinase 2 Inhibitors.
Yogo, T., Nagamiya, H., Seto, M., Sasaki, S., Shih-Chung, H., Ohba, Y., Tokunaga, N., Lee, G.N., Rhim, C.Y., Yoon, C.H., Cho, S.Y., Skene, R., Yamamoto, S., Satou, Y., Kuno, M., Miyazaki, T., Nakagawa, H., Okabe, A., Marui, S., Aso, K., Yoshida, M.(2016) J Med Chem 59: 733-749
- PubMed: 26701356
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01857
- Primary Citation of Related Structures:
5F1Z, 5F20 - PubMed Abstract:
We report herein the discovery and optimization of 3-amino-1,5-dihydro-4H-pyrazolopyridin-4-one TYK2 inhibitors. High-throughput screening against TYK2 and JAK1-3 provided aminoindazole derivative 1 as a hit compound. Scaffold hopping of the aminoindazole core led to the discovery of 3-amino-1,5-dihydro-4H-pyrazolopyridin-4-one derivative 3 as a novel chemotype of TYK2 inhibitors. Interestingly, initial SAR study suggested that this scaffold could have a vertically flipped binding mode, which prompted us to introduce a substituent at the 7-position as a moiety directed toward the solvent-exposed region. Introduction of a 1-methyl-3-pyrazolyl moiety at the 7-position resulted in a dramatic increase in TYK2 inhibitory activity, and further optimization led to the discovery of 20. Compound 20 inhibited IL-23-induced IL-22 production in a rat PD assay, as well as inhibited IL-23 signaling in human PBMC. Furthermore, 20 showed selectivity for IL-23 signaling inhibition against GM-CSF, demonstrating the unique cytokine selectivity of the novel TYK2 inhibitor.
Organizational Affiliation:
Pharmaceutical Research Division, Takeda Pharmaceutical Company Limited , 26-1 Muraoka-Higashi 2-chome, Fujisawa, Kanagawa 251-8555, Japan.