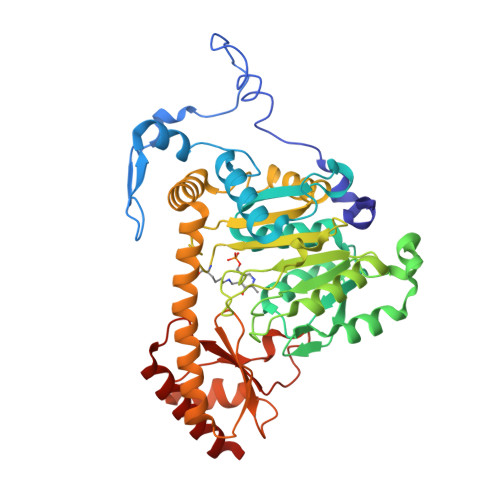
Structure of the PLP-Form of the Human Kynurenine Aminotransferase II in a Novel Spacegroup at 1.83 angstrom Resolution.
Nematollahi, A., Sun, G., Harrop, S.J., Hanrahan, J.R., Church, W.B.(2016) Int J Mol Sci 17: 446-446
- PubMed: 27023527
- DOI: https://doi.org/10.3390/ijms17040446
- Primary Citation of Related Structures:
5EUN - PubMed Abstract:
Kynurenine aminotransferase II (KAT-II) is a 47 kDa pyridoxal phosphate (PLP)-dependent enzyme, active as a homodimer, which catalyses the transamination of the amino acids kynurenine (KYN) and 3-hydroxykynurenine (3-HK) in the tryptophan pathway, and is responsible for producing metabolites that lead to kynurenic acid (KYNA), which is implicated in several neurological diseases such as schizophrenia. In order to fully describe the role of KAT-II in the pathobiology of schizophrenia and other brain disorders, the crystal structure of full-length PLP-form hKAT-II was determined at 1.83 Å resolution, the highest available. The electron density of the active site reveals an aldimine linkage between PLP and Lys263, as well as the active site residues, which characterize the fold-type I PLP-dependent enzymes.
Organizational Affiliation:
Group in Biomolecular Structure and Informatics, Faculty of Pharmacy, University of Sydney, Sydney, NSW 2006, Australia. anem7250@uni.sydney.edu.au.