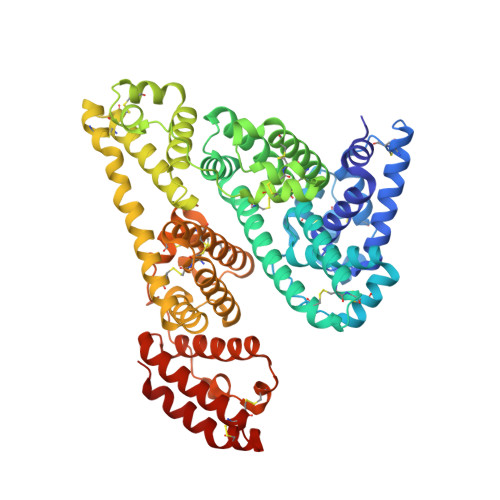
Structural Insights into the Competitive Binding of Diclofenac and Naproxen by Equine Serum Albumin.
Sekula, B., Bujacz, A.(2016) J Med Chem 59: 82-89
- PubMed: 26652101
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00909
- Primary Citation of Related Structures:
4ZBQ, 4ZBR, 5DBY - PubMed Abstract:
The binding modes to equine serum albumin (ESA) of two nonsteroidal anti-inflammatory drugs (NSAIDs), diclofenac (Dic) and naproxen (Nps), were studied by X-ray crystallography and isothermal titration calorimetry. On the basis of the crystal structure of ESA/Dic determined to a resolution of 1.92 Å and the structure of the previously described ESA/Nps complex (2.42 Å), it was found that both NSAIDs bind within drug site 2 (DS2) of ESA and both occupy secondary binding sites in separate cavities of domain II (Nps) and domain III (Dic). The two structures of the ternary complex ESA/Dic/Nps, obtained by competitive cocrystallization (2.19 Å) and through a displacement experiment (2.35 Å), were determined to investigate possible competition of these widely used pharmaceutical drugs in binding to ESA. In these complexes Nps occupies the DS2 pocket common for both drugs, whereas the other distinct binding sites of Dic and Nps remain unaffected. These results suggest that combined application of both drugs may result in increased concentration of free diclofenac in plasma.
Organizational Affiliation:
Institute of Technical Biochemistry, Faculty of Biotechnology and Food Sciences, Lodz University of Technology , Stefanowskiego 4/10, 90-924 Lodz, Poland.