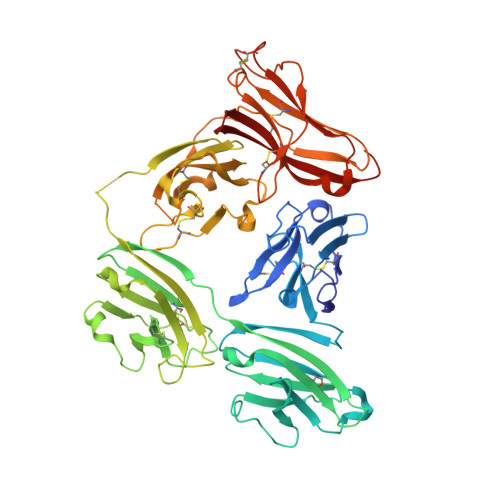
The structure and dynamics of secretory component and its interactions with polymeric immunoglobulins.
Stadtmueller, B.M., Huey-Tubman, K.E., Lopez, C.J., Yang, Z., Hubbell, W.L., Bjorkman, P.J.(2016) Elife 5
- PubMed: 26943617
- DOI: https://doi.org/10.7554/eLife.10640
- Primary Citation of Related Structures:
5D4K, 5F1S - PubMed Abstract:
As a first-line vertebrate immune defense, the polymeric immunoglobulin receptor (pIgR) transports polymeric IgA and IgM across epithelia to mucosal secretions, where the cleaved ectodomain (secretory component; SC) becomes a component of secretory antibodies, or when unliganded, binds and excludes bacteria. Here we report the 2.6Å crystal structure of unliganded human SC (hSC) and comparisons with a 1.7Å structure of teleost fish SC (tSC), an early pIgR ancestor. The hSC structure comprises five immunoglobulin-like domains (D1-D5) arranged as a triangle, with an interface between ligand-binding domains D1 and D5. Electron paramagnetic resonance measurements confirmed the D1-D5 interface in solution and revealed that it breaks upon ligand binding. Together with binding studies of mutant and chimeric SCs, which revealed domain contributions to secretory antibody formation, these results provide detailed models for SC structure, address pIgR evolution, and demonstrate that SC uses multiple conformations to protect mammals from pathogens.
Organizational Affiliation:
Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, United States.