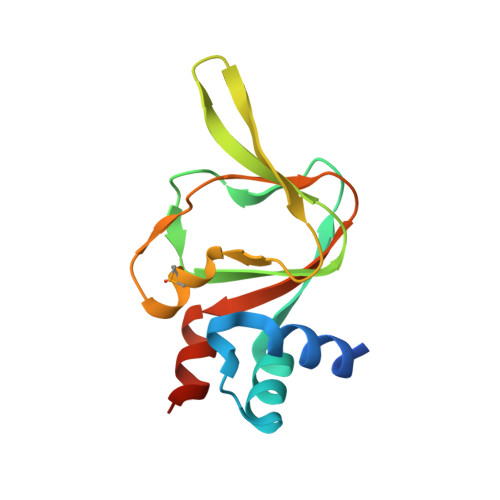
Crystal structure of cyclic nucleotide-binding-like protein from Brucella abortus
He, Z., Gao, Y., Dong, J., Ke, Y., Li, X., Chen, Z., Zhang, X.C.(2015) Biochem Biophys Res Commun 468: 647-652
- PubMed: 26549229
- DOI: https://doi.org/10.1016/j.bbrc.2015.11.005
- Primary Citation of Related Structures:
5D1I - PubMed Abstract:
The cyclic nucleotide-binding (CNB)-like protein (CNB-L) from Brucella abortus shares sequence homology with CNB domain-containing proteins. We determined the crystal structure of CNB-L at 2.0 Å resolution in the absence of its C-terminal helix and nucleotide. The 3D structure of CNB-L is in a two-fold symmetric form. Each protomer shows high structure similarity to that of cGMP-binding domain-containing proteins, and likely mimics their nucleotide-free conformation. A key residue, Glu17, mediates the dimerization and prevents binding of cNMP to the canonical ligand-pocket. The structurally observed dimer of CNB-L is stable in solution, and thus is likely to be biologically relevant.
Organizational Affiliation:
National Laboratory of Macromolecules, National Center of Protein Science- Beijing, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Beijing, 100101, China; University of Chinese Academy of Sciences, Beijing, 100049, China.