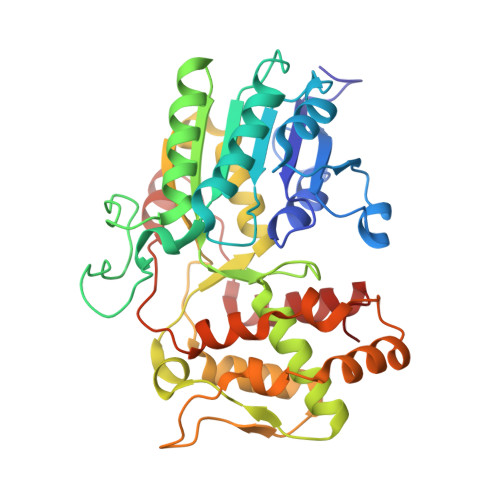
Structure of iridoid synthase in complex with NADP(+)/8-oxogeranial reveals the structural basis of its substrate specificity.
Qin, L., Zhu, Y., Ding, Z., Zhang, X., Ye, S., Zhang, R.(2016) J Struct Biol 194: 224-230
- PubMed: 26868105
- DOI: https://doi.org/10.1016/j.jsb.2016.02.010
- Primary Citation of Related Structures:
5COA, 5COB - PubMed Abstract:
Iridoid synthase (IS), as a vegetal enzyme belonging to the short-chain dehydrogenase/reductase (SDR) superfamily, produces the ring skeletons for downstream alkaloids with various pharmaceutical activities, including the commercially available antineoplastic agents, vinblastine and vincristine. Here, we present the crystal structures of IS in apo state and in complex with NADP(+)/8-oxogeranial, exhibiting an active center that lacks the classical Tyr/Lys/Ser triad spatially conserved in SDRs, with only the catalytically critical function of triad tyrosine remained in Tyr178. In consistent, mutation of Tyr178 to a phenylalanine residue significantly abolished the catalytic activity of IS. Within the substrate binding pocket, the linear-shaped 8-oxogeranial adopts an entirely extended conformation with its two aldehyde ends hydrogen-bonded to Tyr178-OH and Ser349-OH, respectively. In addition, the intermediate carbon chain of bound substrate is harbored by a well-ordered hydrophobic scaffold, involving residues Ile145, Phe149, Leu203, Met213, Phe342, Ile345 and Leu352. Mutagenesis studies showed that both Ser349 and the hydrophobic residues around are determinant to the substrate specificity and, consequently, the catalytic activity of IS. In contrast, the Gly150-Pro160 loop previously proposed as a factor involved in substrate binding might have very limited contribution, because the deletion of residues Ile151-His161 has only slight influence on the catalytic activity. We believe that the present work will help to elucidate the substrate specificity of IS and to integrate its detailed catalytic mechanism.
Organizational Affiliation:
National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, People's Republic of China; University of Chinese Academy of Sciences, Beijing 100049, People's Republic of China.