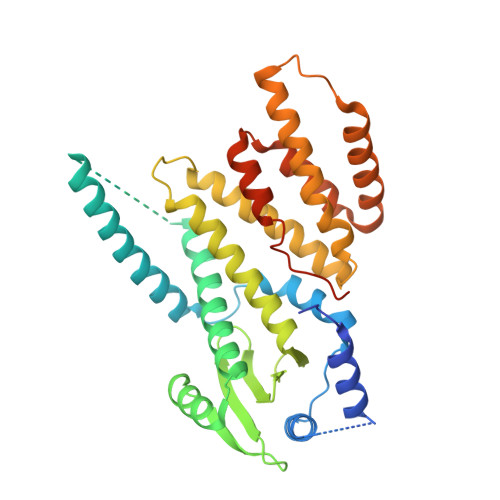
Structure and Function of the N-Terminal Domain of the Vesicular Stomatitis Virus RNA Polymerase.
Qiu, S., Ogino, M., Luo, M., Ogino, T., Green, T.J.(2015) J Virol 90: 715-724
- PubMed: 26512087
- DOI: https://doi.org/10.1128/JVI.02317-15
- Primary Citation of Related Structures:
5CHS - PubMed Abstract:
Viruses have various mechanisms to duplicate their genomes and produce virus-specific mRNAs. Negative-strand RNA viruses encode their own polymerases to perform each of these processes. For the nonsegmented negative-strand RNA viruses, the polymerase is comprised of the large polymerase subunit (L) and the phosphoprotein (P). L proteins from members of the Rhabdoviridae, Paramyxoviridae, and Filoviridae share sequence and predicted secondary structure homology. Here, we present the structure of the N-terminal domain (conserved region I) of the L protein from a rhabdovirus, vesicular stomatitis virus, at 1.8-Å resolution. The strictly and strongly conserved residues in this domain cluster in a single area of the protein. Serial mutation of these residues shows that many of the amino acids are essential for viral transcription but not for mRNA capping. Three-dimensional alignments show that this domain shares structural homology with polymerases from other viral families, including segmented negative-strand RNA and double-stranded RNA (dsRNA) viruses.
Organizational Affiliation:
Department of Microbiology, School of Medicine, University of Alabama at Birmingham, Birmingham, Alabama, USA.