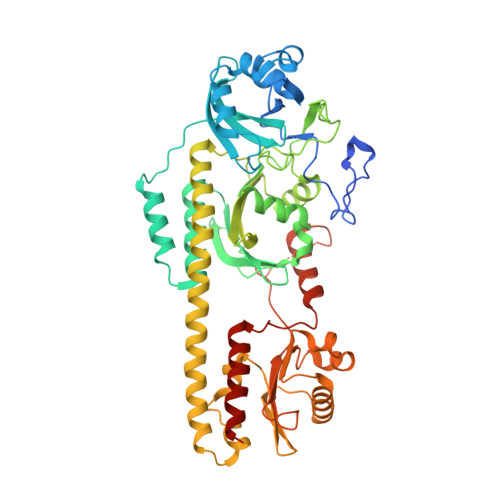
Crystal Structure of Deinococcus Phytochrome in the Photoactivated State Reveals a Cascade of Structural Rearrangements during Photoconversion.
Burgie, E.S., Zhang, J., Vierstra, R.D.(2016) Structure 24: 448-457
- PubMed: 26853942
- DOI: https://doi.org/10.1016/j.str.2016.01.001
- Primary Citation of Related Structures:
5C5K - PubMed Abstract:
Phytochromes are photochromic photoreceptors responsible for a myriad of red/far-red light-dependent processes in plants and microorganisms. Interconversion is initially driven by photoreversible isomerization of bilin, but how this alteration directs the photostate-dependent changes within the protein to actuate signaling is poorly understood. Here, we describe the structure of the Deinococcus phytochrome photosensory module in its near complete far-red light-absorbing Pfr state. In addition to confirming the 180° rotation of the D-pyrrole ring, the dimeric structure clearly identifies downstream rearrangements that trigger large-scale conformational differences between the dark-adapted and photoactivated states. Mutational analyses verified the importance of residues surrounding the bilin in Pfr stabilization, and protease sensitivity assays corroborated photostate alterations that propagate along the dimeric interface. Collectively, these data support a cooperative "toggle" model for phytochrome photoconversion and advance our understanding of the allosteric connection between the photosensory and output modules.
Organizational Affiliation:
Department of Genetics, University of Wisconsin-Madison, 425-G Henry Mall, Madison, WI 53706, USA; Department of Biology, Washington University in St Louis, Campus Box 1137, One Brookings Drive, St. Louis, MO 63130, USA.