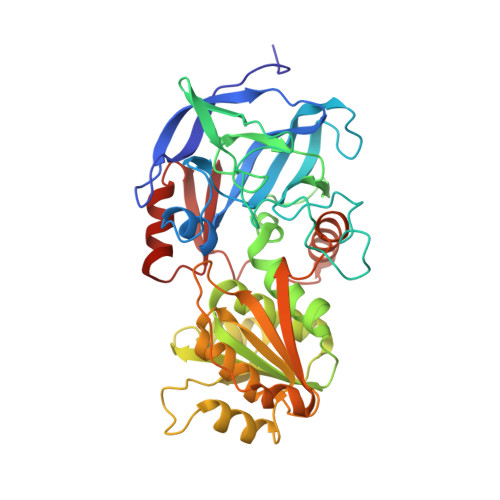
Interdomain motion in liver alcohol dehydrogenase. Structural and energetic analysis of the hinge bending mode.
Colonna-Cesari, F., Perahia, D., Karplus, M., Eklund, H., Braden, C.I., Tapia, O.(1986) J Biol Chem 261: 15273-15280
- PubMed: 3771574
- Primary Citation of Related Structures:
5ADH, 8ADH - PubMed Abstract:
A study of the hinge bending mode in the enzyme liver alcohol dehydrogenase is made by use of empirical energy functions. The enzyme is a dimer, with each monomer composed of a coenzyme binding domain and a catalytic domain with a large cleft between the two. Superposition of the apoenzyme and holoenzyme crystal structures is used to determine a rigid rotation axis for closing of the cleft. It is shown that a rigid body transformation of the apoenzyme to the holoenzyme structure corresponds to a 10 degrees rotation of the catalytic domain about this axis. The rotation is not along the least-motion path for closing of the cleft but instead corresponds to the catalytic domain coming closer to the coenzyme binding domain by a sliding motion. Estimation of the energy associated with the interdomain motion of the apoenzyme over a range of 90 degrees (-40 to 50 degrees, where 0 degrees corresponds to the minimized crystal structure) demonstrates that local structural relaxation makes possible large-scale rotations with relatively small energy increments. A variety of structural rearrangements associated with the domain motion are characterized. They involve the hinge region residues that provide the covalent connections between the two domains and certain loop regions that are brought into contact by the rotation. Differences between the energy minimized and the holoenzyme structures point to the existence of alternative conformations for loops and to the importance of the ligands in the structural rearrangements.