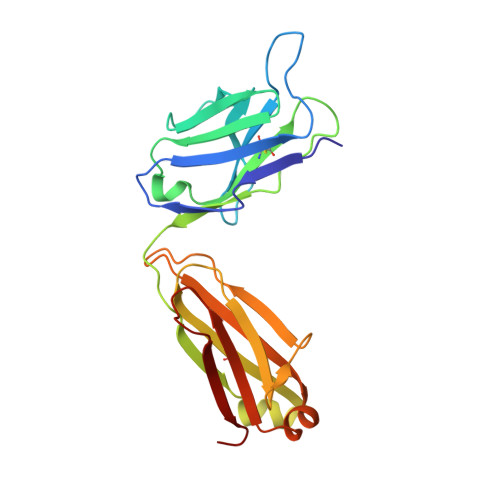
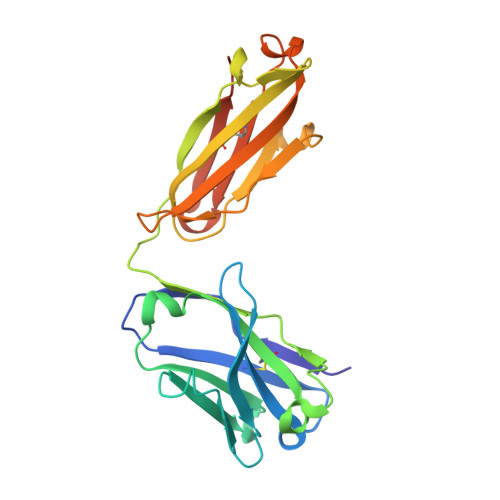
Structural insights into humanization of anti-tissue factor antibody 10H10.
Teplyakov, A., Obmolova, G., Malia, T.J., Raghunathan, G., Martinez, C., Fransson, J., Edwards, W., Connor, J., Husovsky, M., Beck, H., Chi, E., Fenton, S., Zhou, H., Almagro, J.C., Gilliland, G.L.(2018) MAbs 10: 269-277
- PubMed: 29283291
- DOI: https://doi.org/10.1080/19420862.2017.1412026
- Primary Citation of Related Structures:
5W05, 5W06 - PubMed Abstract:
Murine antibody 10H10 raised against human tissue factor is unique in that it blocks the signaling pathway, and thus inhibits angiogenesis and tumor growth without interfering with coagulation. As a potential therapeutic, the antibody was humanized in a two-step procedure. Antigen-binding loops were grafted onto selected human frameworks and the resulting chimeric antibody was subjected to affinity maturation by using phage display libraries. The results of humanization were analyzed from the structural perspective through comparison of the structure of a humanized variant with the parental mouse antibody. This analysis revealed several hot spots in the framework region that appear to affect antigen binding, and therefore should be considered in human germline selection. In addition, some positions in the Vernier zone, e.g., residue 71 in the heavy chain, that are traditionally thought to be crucial appear to tolerate amino acid substitutions without any effect on binding. Several humanized variants were produced using both short and long forms of complementarity-determining region (CDR) H2 following the difference in the Kabat and Martin definitions. Comparison of such pairs indicated consistently higher thermostability of the variants with short CDR H2. Analysis of the binding data in relation to the structures singled out the ImMunoGeneTics information system® germline IGHV1-2*01 as dubious owing to two potentially destabilizing mutations as compared to the other alleles of the same germline and to other human germlines.
Organizational Affiliation:
a Janssen Research and Development, LLC , 1400 McKean Road, Spring House, PA , USA.