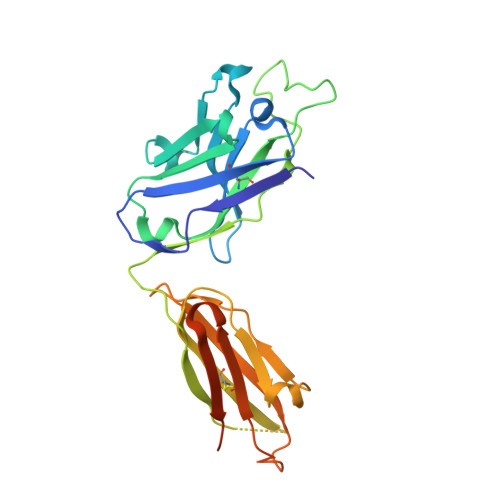
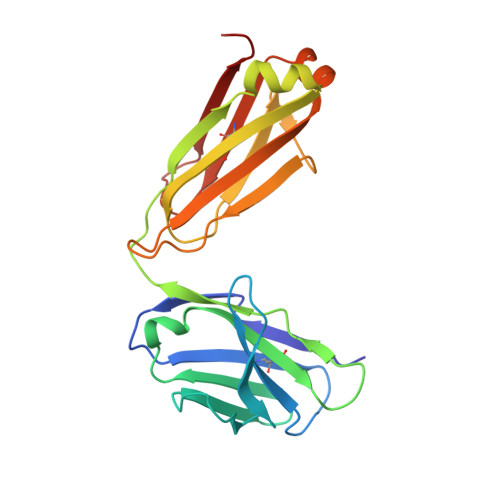
Asymmetric recognition of HIV-1 Envelope trimer by V1V2 loop-targeting antibodies.
Wang, H., Gristick, H.B., Scharf, L., West, A.P., Galimidi, R.P., Seaman, M.S., Freund, N.T., Nussenzweig, M.C., Bjorkman, P.J.(2017) Elife 6
- PubMed: 28548638
- DOI: https://doi.org/10.7554/eLife.27389
- Primary Citation of Related Structures:
5VIY, 5VJ6, 5VVF - PubMed Abstract:
The HIV-1 envelope (Env) glycoprotein binds to host cell receptors to mediate membrane fusion. The prefusion Env trimer is stabilized by V1V2 loops that interact at the trimer apex. Broadly neutralizing antibodies (bNAbs) against V1V2 loops, exemplified by PG9, bind asymmetrically as a single Fab to the apex of the symmetric Env trimer using a protruding CDRH3 to penetrate the Env glycan shield. Here we characterized a distinct mode of V1V2 epitope recognition by the new bNAb BG1 in which two Fabs bind asymmetrically per Env trimer using a compact CDRH3. Comparisons between cryo-EM structures of Env trimer complexed with BG1 (6.2 Å resolution) and PG9 (11.5 Å resolution) revealed a new V1V2-targeting strategy by BG1. Analyses of the EM structures provided information relevant to vaccine design including molecular details for different modes of asymmetric recognition of Env trimer and a binding model for BG1 recognition of V1V2 involving glycan flexibility.
Organizational Affiliation:
Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, United States.