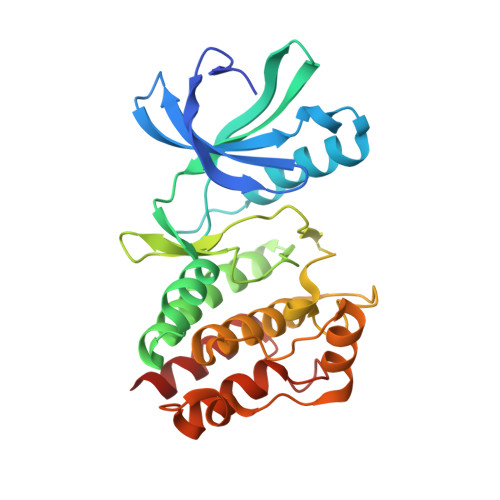
In Silico Screen and Structural Analysis Identifies Bacterial Kinase Inhibitors which Act with beta-Lactams To Inhibit Mycobacterial Growth.
Wlodarchak, N., Teachout, N., Beczkiewicz, J., Procknow, R., Schaenzer, A.J., Satyshur, K., Pavelka, M., Zuercher, W., Drewry, D., Sauer, J.D., Striker, R.(2018) Mol Pharm 15: 5410-5426
- PubMed: 30285456
- DOI: https://doi.org/10.1021/acs.molpharmaceut.8b00905
- Primary Citation of Related Structures:
5U94 - PubMed Abstract:
New tools and concepts are needed to combat antimicrobial resistance. Actinomycetes and firmicutes share several eukaryotic-like Ser/Thr kinases (eSTK) that offer antibiotic development opportunities, including PknB, an essential mycobacterial eSTK. Despite successful development of potent biochemical PknB inhibitors by many groups, clinically useful microbiologic activity has been elusive. Additionally, PknB kinetics are not fully described, nor are structures with specific inhibitors available to inform inhibitor design. We used computational modeling with available structural information to identify human kinase inhibitors predicted to bind PknB, and we selected hits based on drug-like characteristics intended to increase the likelihood of cell entry. The computational model suggested a family of inhibitors, the imidazopyridine aminofurazans (IPAs), bind PknB with high affinity. We performed an in-depth characterization of PknB and found that these inhibitors biochemically inhibit PknB, with potency roughly following the predicted models. A novel X-ray structure confirmed that the inhibitors bound as predicted and made favorable protein contacts with the target. These inhibitors also have antimicrobial activity toward mycobacteria and nocardia. We demonstrated that the inhibitors are uniquely potentiated by β-lactams but not antibiotics traditionally used to treat mycobacteria, consistent with PknB's role in sensing cell wall stress. This is the first demonstration in the phylum actinobacteria that some β-lactam antibiotics could be more effective if paired with a PknB inhibitor. Collectively, our data show that in silico modeling can be used as a tool to discover promising drug leads, and the inhibitors we discovered can act with clinically relevant antibiotics to restore their efficacy against bacteria with limited treatment options.
Organizational Affiliation:
Department of Medicine , University of Wisconsin-Madison , 3341 Microbial Sciences Building, 1550 Linden Dr. , Madison , Wisconsin 53706 , United States.