5JNX
The 6.6 A cryo-EM structure of the full-length human NPC1 in complex with the cleaved glycoprotein of Ebola virus
- PDB DOI: https://doi.org/10.2210/pdb5JNX/pdb
- EM Map EMD-8169: EMDB EMDataResource
- Classification: MEMBRANE PROTEIN
- Organism(s): Homo sapiens, Ebola virus - Zaire (1995)
- Expression System: Homo sapiens, Insect cell expression vector pTIE1
- Mutation(s): Yes
- Membrane Protein: Yes PDBTMmpstruc
- Deposited: 2016-05-01 Released: 2016-06-15
- Funding Organization(s): Ministry of Science and Technology of China, National Natural Science Foundation of China, President Foundation of Chinese Academy of Sciences and Strategic Priority Research Program of the Chinese Academy of Sciences
Experimental Data Snapshot
- Method: ELECTRON MICROSCOPY
- Resolution: 6.56 Å
- Aggregation State: PARTICLE
- Reconstruction Method: SINGLE PARTICLE
wwPDB Validation 3D Report Full Report
This is version 2.1 of the entry. See complete history.
Macromolecules
Find similar proteins by:
(by identity cutoff) | 3D Structure
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Niemann-Pick C1 protein | 1,278 | Homo sapiens | Mutation(s): 0 Gene Names: NPC1 Membrane Entity: Yes | 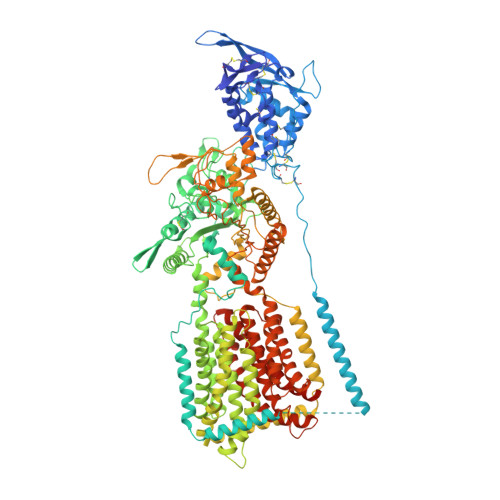 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for O15118 (Homo sapiens) Explore O15118 Go to UniProtKB: O15118 | |||||
PHAROS: O15118 GTEx: ENSG00000141458 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O15118 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by:
(by identity cutoff) | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Envelope glycoprotein | B [auth C], D [auth E], F [auth G] | 158 | Ebola virus - Zaire (1995) | Mutation(s): 1 Gene Names: GP Membrane Entity: Yes | 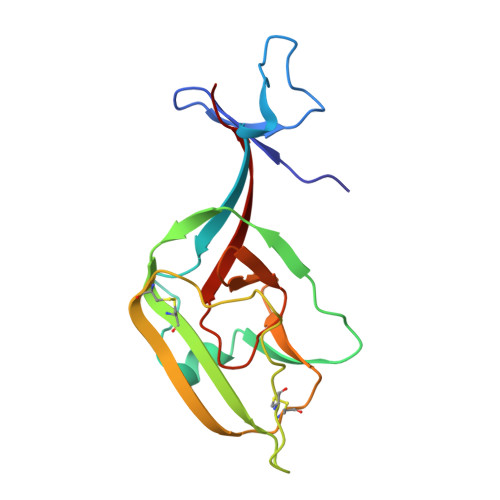 |
UniProt | |||||
Find proteins for P87666 (Zaire ebolavirus (strain Kikwit-95)) Explore P87666 Go to UniProtKB: P87666 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P87666 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by:
(by identity cutoff) | 3D Structure
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Envelope glycoprotein | C [auth D], E [auth F], G [auth H] | 130 | Ebola virus - Zaire (1995) | Mutation(s): 0 Gene Names: GP Membrane Entity: Yes |  |
UniProt | |||||
Find proteins for P87666 (Zaire ebolavirus (strain Kikwit-95)) Explore P87666 Go to UniProtKB: P87666 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P87666 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Oligosaccharides
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | H [auth B], I, K, M, N, H [auth B], I, K, M, N, O, P | 2 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G42666HT GlyCosmos: G42666HT GlyGen: G42666HT | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | J | 3 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G15407YE GlyCosmos: G15407YE GlyGen: G15407YE | |||||
Entity ID: 6 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| alpha-D-mannopyranose-(1-6)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | L | 4 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G22573RC GlyCosmos: G22573RC GlyGen: G22573RC | |||||
Small Molecules
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Query on NAG | Q [auth A] R [auth A] S [auth A] T [auth A] U [auth A] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
Experimental Data & Validation
Experimental Data
- Method: ELECTRON MICROSCOPY
- Resolution: 6.56 Å
- Aggregation State: PARTICLE
- Reconstruction Method: SINGLE PARTICLE
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | RELION | 1.4 |
Entry History & Funding Information
Deposition Data
- Released Date: 2016-06-15 Deposition Author(s): Gong, X., Qian, H.W., Zhou, X.H., Wu, J.P., Wan, T., Shi, Y., Gao, F., Zhou, Q., Yan, N.
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ministry of Science and Technology of China | China | 2015CB9101012014, ZX09507003006 |
| National Natural Science Foundation of China | China | project 31321062 and 81590761 |
| President Foundation of Chinese Academy of Sciences and Strategic Priority Research Program of the Chinese Academy of Sciences | China | XDB08020100 |
Revision History (Full details and data files)
- Version 1.0: 2016-06-15
Type: Initial release - Version 1.1: 2017-12-13
Changes: Data processing, Database references - Version 1.2: 2019-10-23
Changes: Advisory, Data collection, Database references, Derived calculations - Version 1.3: 2019-12-04
Changes: Data collection - Version 1.4: 2019-12-11
Changes: Data collection - Version 2.0: 2020-07-29
Type: Remediation
Reason: Carbohydrate remediation
Changes: Advisory, Atomic model, Data collection, Derived calculations, Structure summary - Version 2.1: 2020-08-05
Changes: Derived calculations, Source and taxonomy, Structure summary













