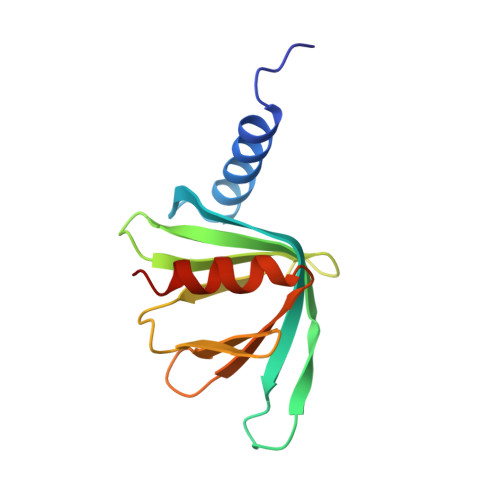
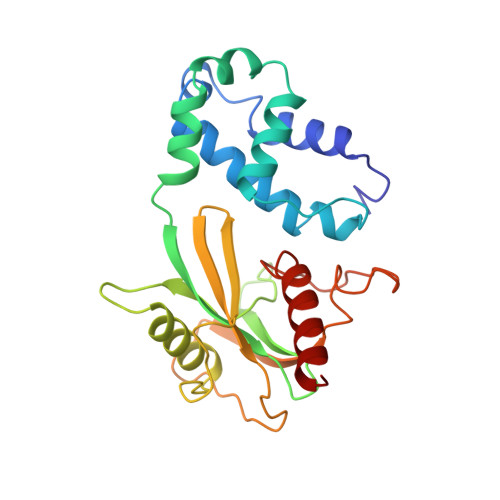
Structure of the Dcp2-Dcp1 mRNA-decapping complex in the activated conformation.
Valkov, E., Muthukumar, S., Chang, C.T., Jonas, S., Weichenrieder, O., Izaurralde, E.(2016) Nat Struct Mol Biol 23: 574-579
- PubMed: 27183195
- DOI: https://doi.org/10.1038/nsmb.3232
- Primary Citation of Related Structures:
5J3Q, 5J3T, 5J3Y - PubMed Abstract:
The removal of the mRNA 5' cap (decapping) by Dcp2 shuts down translation and commits mRNA to full degradation. Dcp2 activity is enhanced by activator proteins such as Dcp1 and Edc1. However, owing to conformational flexibility, the active conformation of Dcp2 and the mechanism of decapping activation have remained unknown. Here, we report a 1.6-Å-resolution crystal structure of the Schizosaccharomyces pombe Dcp2-Dcp1 heterodimer in an unprecedented conformation that is tied together by an intrinsically disordered peptide from Edc1. In this ternary complex, an unforeseen rotation of the Dcp2 catalytic domain allows residues from both Dcp2 and Dcp1 to cooperate in RNA binding, thus explaining decapping activation by increased substrate affinity. The architecture of the Dcp2-Dcp1-Edc1 complex provides a rationale for the conservation of a sequence motif in Edc1 that is also present in unrelated decapping activators, thus indicating that the presently described mechanism of decapping activation is evolutionarily conserved.
Organizational Affiliation:
Department of Biochemistry, Max Planck Institute for Developmental Biology, Tübingen, Germany.