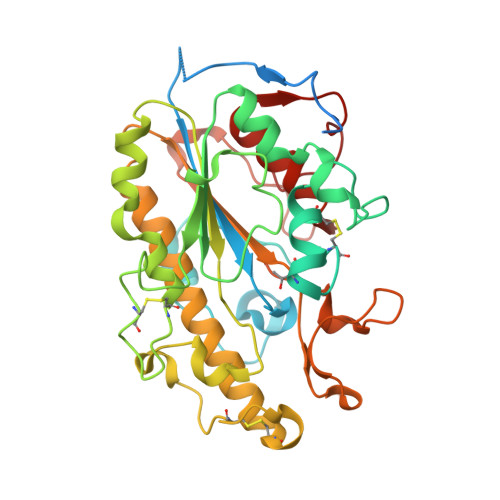
Crystal Structure of Escherichia coli originated MCR-1, a phosphoethanolamine transferase for Colistin Resistance.
Hu, M., Guo, J., Cheng, Q., Yang, Z., Chan, E.W.C., Chen, S., Hao, Q.(2016) Sci Rep 6: 38793-38793
- PubMed: 27958270
- DOI: https://doi.org/10.1038/srep38793
- PubMed Abstract:
MCR-1 is a phosphoethanolamine (pEtN) transferase that modifies the pEtN moiety of lipid A, conferring resistance to colistin, which is an antibiotic belonging to the class of polypeptide antibiotics known as polymyxins and is the last-line antibiotic used to treat multidrug resistant bacterial infections. Here we determined the crystal structure of the catalytic domain of MCR-1 (MCR-1-ED), which is originated in Escherichia coli (E. coli). MCR-1-ED was found to comprise several classical β-α-β-α motifs that constitute a "sandwich" conformation. Two interlaced molecules with different phosphorylation status of the residue T285 could give rise to two functional statuses of MCR-1 depending on the physiological conditions. MCR-1, like other known pEtN transferases, possesses an enzymatic site equipped with zinc binding residues. Interestingly, two zinc ions were found to mediate intermolecular interactions between MCR-1-ED molecules in one asymmetric unit and hence concatenation of MCR-1, allowing the protein to be oligomer. Findings of this work shall provide important insight into development of effective and clinically useful inhibitors of MCR-1 or structurally similar enzymes.
Organizational Affiliation:
School of Biomedical Sciences, University of Hong Kong, Laboratory Block, 21 Sassoon Road, Pokfulam, Hong Kong, China.