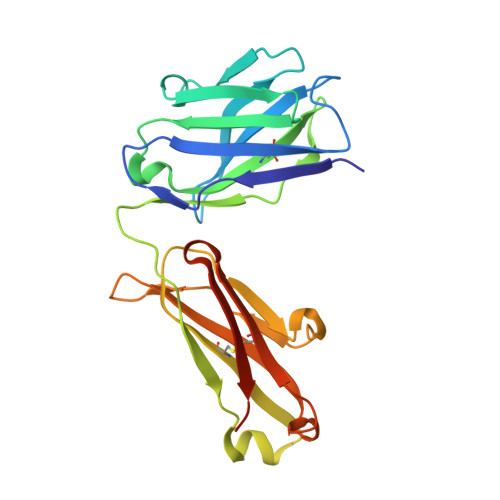
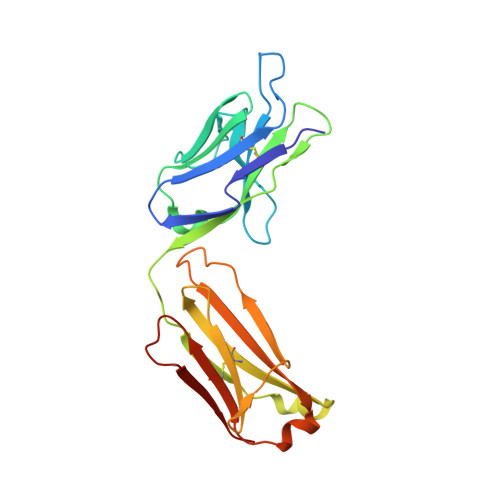
MoFvAb: Modeling the Fv region of antibodies.
Bujotzek, A., Fuchs, A., Qu, C., Benz, J., Klostermann, S., Antes, I., Georges, G.(2015) MAbs 7: 838-852
- PubMed: 26176812
- DOI: https://doi.org/10.1080/19420862.2015.1068492
- Primary Citation of Related Structures:
5BMF - PubMed Abstract:
Knowledge of the 3-dimensional structure of the antigen-binding region of antibodies enables numerous useful applications regarding the design and development of antibody-based drugs. We present a knowledge-based antibody structure prediction methodology that incorporates concepts that have arisen from an applied antibody engineering environment. The protocol exploits the rich and continuously growing supply of experimentally derived antibody structures available to predict CDR loop conformations and the packing of heavy and light chain quickly and without user intervention. The homology models are refined by a novel antibody-specific approach to adapt and rearrange sidechains based on their chemical environment. The method achieves very competitive all-atom root mean square deviation values in the order of 1.5 Å on different evaluation datasets consisting of both known and previously unpublished antibody crystal structures.
Organizational Affiliation:
a Roche Pharmaceutical Research and Early Development; Large Molecule Research; Roche Innovation Center Penzberg ; Penzberg , Germany.