The crystal structure of mupain-1-IG in complex with murinised human uPA at pH7.4
Jiang, L., Andreasen, P.A., Huang, M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Urokinase-type plasminogen activator | A [auth U] | 247 | Homo sapiens | Mutation(s): 3 Gene Names: PLAU EC: 3.4.21.73 | 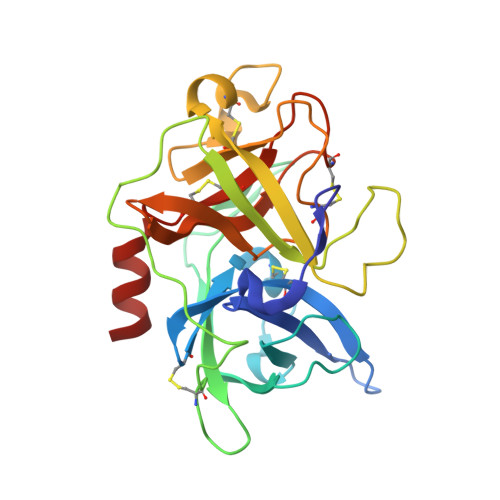 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P00749 (Homo sapiens) Explore P00749 Go to UniProtKB: P00749 | |||||
PHAROS: P00749 GTEx: ENSG00000122861 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00749 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| upain-1-W3A | B [auth P] | 12 | synthetic construct | Mutation(s): 0 | 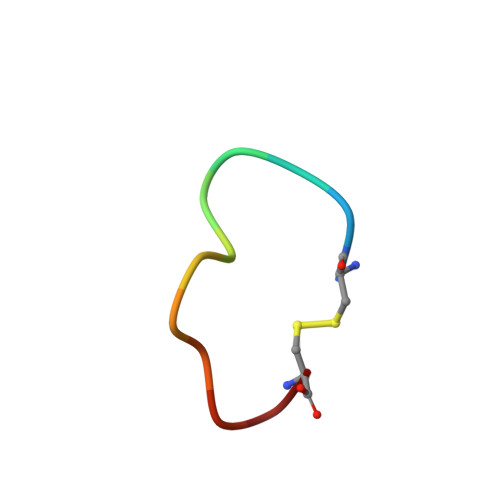 |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 120.751 | α = 90 |
| b = 120.751 | β = 90 |
| c = 42.961 | γ = 120 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data scaling |
| Coot | model building |